[English] 日本語
 Yorodumi
Yorodumi- PDB-7y48: Cryo-EM Structure of biliverdin-bound mitochondrial ABC transport... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 7y48 | ||||||
|---|---|---|---|---|---|---|---|
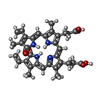
| Title | Cryo-EM Structure of biliverdin-bound mitochondrial ABC transporter ABCB10 from Biortus | ||||||
 Components Components | ATP-binding cassette sub-family B member 10, mitochondrial | ||||||
 Keywords Keywords | MEMBRANE PROTEIN / ABC transporter / biliverdin | ||||||
| Function / homology |  Function and homology information Function and homology informationexport from the mitochondrion / positive regulation of heme biosynthetic process / Mitochondrial ABC transporters / positive regulation of hemoglobin biosynthetic process / mitochondrial unfolded protein response / Translocases; Catalysing the translocation of other compounds; Linked to the hydrolysis of a nucleoside triphosphate / heme biosynthetic process / mitochondrial transport / erythrocyte development / ABC-type transporter activity ...export from the mitochondrion / positive regulation of heme biosynthetic process / Mitochondrial ABC transporters / positive regulation of hemoglobin biosynthetic process / mitochondrial unfolded protein response / Translocases; Catalysing the translocation of other compounds; Linked to the hydrolysis of a nucleoside triphosphate / heme biosynthetic process / mitochondrial transport / erythrocyte development / ABC-type transporter activity / positive regulation of erythrocyte differentiation / mitochondrial membrane / mitochondrial inner membrane / protein homodimerization activity / ATP hydrolysis activity / mitochondrion / ATP binding / metal ion binding Similarity search - Function | ||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||
| Method | ELECTRON MICROSCOPY / single particle reconstruction / cryo EM / Resolution: 2.85 Å | ||||||
 Authors Authors | Cao, S. / Yang, Y. | ||||||
| Funding support | 1items
| ||||||
 Citation Citation |  Journal: Nat Commun / Year: 2023 Journal: Nat Commun / Year: 2023Title: Cryo-EM structures of mitochondrial ABC transporter ABCB10 in apo and biliverdin-bound form. Authors: Sheng Cao / Yihu Yang / Lili He / Yumo Hang / Xiaodong Yan / Hui Shi / Jiaquan Wu / Zhuqing Ouyang /  Abstract: ABCB10, a member of ABC transporter superfamily that locates in the inner membrane of mitochondria, plays crucial roles in hemoglobin synthesis, antioxidative stress and stabilization of the iron ...ABCB10, a member of ABC transporter superfamily that locates in the inner membrane of mitochondria, plays crucial roles in hemoglobin synthesis, antioxidative stress and stabilization of the iron transporter mitoferrin-1. Recently, it was found that ABCB10 is a mitochondrial biliverdin exporter. However, the molecular mechanism of biliverdin export by ABCB10 remains elusive. Here we report the cryo-EM structures of ABCB10 in apo (ABCB10-apo) and biliverdin-bound form (ABCB10-BV) at 3.67 Å and 2.85 Å resolution, respectively. ABCB10-apo adopts a wide-open conformation and may thus represent the apo form structure. ABCB10-BV forms a closed conformation and biliverdin situates in a hydrophobic pocket in one protomer and bridges the interaction through hydrogen bonds with the opposing one. We also identify cholesterols sandwiched by BVs and discuss the export dynamics based on these structural and biochemical observations. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  7y48.cif.gz 7y48.cif.gz | 141.6 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb7y48.ent.gz pdb7y48.ent.gz | 102.2 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  7y48.json.gz 7y48.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/y4/7y48 https://data.pdbj.org/pub/pdb/validation_reports/y4/7y48 ftp://data.pdbj.org/pub/pdb/validation_reports/y4/7y48 ftp://data.pdbj.org/pub/pdb/validation_reports/y4/7y48 | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  33603MC  7y49C M: map data used to model this data C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
|
|---|---|
| 1 |
|
- Components
Components
| #1: Protein | Mass: 67180.125 Da / Num. of mol.: 1 / Mutation: Q9NRK6 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Gene: ABCB10 / Cell line (production host): HEK293 / Production host: Homo sapiens (human) / Gene: ABCB10 / Cell line (production host): HEK293 / Production host:  Homo sapiens (human) / References: UniProt: Q9NRK6 Homo sapiens (human) / References: UniProt: Q9NRK6 |
|---|---|
| #2: Chemical | ChemComp-CLR / |
| #3: Chemical | ChemComp-IE5 / |
| Has ligand of interest | Y |
-Experimental details
-Experiment
| Experiment | Method: ELECTRON MICROSCOPY |
|---|---|
| EM experiment | Aggregation state: PARTICLE / 3D reconstruction method: single particle reconstruction |
- Sample preparation
Sample preparation
| Component | Name: Complex of ABCB10 binding with biliverdin / Type: COMPLEX / Entity ID: #1 / Source: RECOMBINANT |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Source (recombinant) | Organism:  Homo sapiens (human) / Cell: HEK293 Homo sapiens (human) / Cell: HEK293 |
| Buffer solution | pH: 7.5 |
| Specimen | Conc.: 4.78 mg/ml / Embedding applied: NO / Shadowing applied: NO / Staining applied: NO / Vitrification applied: YES |
| Specimen support | Details: 5W / Grid material: COPPER / Grid mesh size: 300 divisions/in. / Grid type: Quantifoil R1.2/1.3 |
| Vitrification | Instrument: FEI VITROBOT MARK IV / Cryogen name: ETHANE / Humidity: 100 % / Chamber temperature: 277 K |
- Electron microscopy imaging
Electron microscopy imaging
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
|---|---|
| Microscopy | Model: TFS KRIOS |
| Electron gun | Electron source:  FIELD EMISSION GUN / Accelerating voltage: 300 kV / Illumination mode: FLOOD BEAM FIELD EMISSION GUN / Accelerating voltage: 300 kV / Illumination mode: FLOOD BEAM |
| Electron lens | Mode: BRIGHT FIELD / Nominal defocus max: 1800 nm / Nominal defocus min: 1100 nm / Cs: 2.7 mm / C2 aperture diameter: 50 µm |
| Image recording | Average exposure time: 2.5 sec. / Electron dose: 55.21 e/Å2 / Film or detector model: GATAN K3 (6k x 4k) |
- Processing
Processing
| CTF correction | Type: PHASE FLIPPING AND AMPLITUDE CORRECTION |
|---|---|
| Symmetry | Point symmetry: C2 (2 fold cyclic) |
| 3D reconstruction | Resolution: 2.85 Å / Resolution method: FSC 0.143 CUT-OFF / Num. of particles: 206922 / Symmetry type: POINT |
| Atomic model building | B value: 97.3 / Protocol: AB INITIO MODEL |
| Atomic model building | PDB-ID: 4AYX Accession code: 4AYX / Source name: PDB / Type: experimental model |
 Movie
Movie Controller
Controller







 PDBj
PDBj