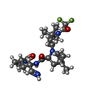
Entry Database : PDB / ID : 7vh8Title Crystal structure of SARS-CoV-2 main protease in complex with protease inhibitor PF-07321332 3C-like proteinase Keywords / / / Function / homology Function Domain/homology Component
/ / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / Biological species Method / / / Resolution : 1.59 Å Authors Zhao, Y. / Zhang, Q. / Yang, H. / Rao, Z. Funding support Organization Grant number Country National Natural Science Foundation of China (NSFC)
Journal : Protein Cell / Year : 2022Title : Crystal structure of SARS-CoV-2 main protease in complex with protease inhibitor PF-07321332.Authors : Zhao, Y. / Fang, C. / Zhang, Q. / Zhang, R. / Zhao, X. / Duan, Y. / Wang, H. / Zhu, Y. / Feng, L. / Zhao, J. / Shao, M. / Yang, X. / Zhang, L. / Peng, C. / Yang, K. / Ma, D. / Rao, Z. / Yang, H. History Deposition Sep 21, 2021 Deposition site / Processing site Revision 1.0 Nov 3, 2021 Provider / Type Revision 2.0 Dec 22, 2021 Group Atomic model / Author supporting evidence ... Atomic model / Author supporting evidence / Data collection / Derived calculations / Non-polymer description / Structure summary Category atom_site / chem_comp ... atom_site / chem_comp / entity / pdbx_entity_instance_feature / pdbx_entity_nonpoly / pdbx_nonpoly_scheme / struct_conn Item _atom_site.B_iso_or_equiv / _atom_site.Cartn_x ... _atom_site.B_iso_or_equiv / _atom_site.Cartn_x / _atom_site.Cartn_y / _atom_site.Cartn_z / _atom_site.auth_atom_id / _atom_site.auth_comp_id / _atom_site.label_atom_id / _atom_site.label_comp_id / _atom_site.type_symbol / _chem_comp.formula / _chem_comp.formula_weight / _chem_comp.id / _chem_comp.name / _chem_comp.pdbx_synonyms / _chem_comp.type / _entity.formula_weight / _entity.pdbx_description / _pdbx_entity_instance_feature.auth_comp_id / _pdbx_entity_instance_feature.comp_id / _pdbx_entity_nonpoly.comp_id / _pdbx_entity_nonpoly.name / _pdbx_nonpoly_scheme.mon_id / _pdbx_nonpoly_scheme.pdb_mon_id / _struct_conn.ptnr2_auth_comp_id / _struct_conn.ptnr2_label_atom_id / _struct_conn.ptnr2_label_comp_id Revision 2.1 Jul 6, 2022 Group / Category Item _citation.journal_volume / _citation.page_first ... _citation.journal_volume / _citation.page_first / _citation.page_last / _citation.year Revision 3.0 Jul 5, 2023 Group Atomic model / Author supporting evidence ... Atomic model / Author supporting evidence / Data collection / Derived calculations / Non-polymer description / Refinement description / Source and taxonomy / Structure summary Category atom_site / chem_comp ... atom_site / chem_comp / entity / entity_src_gen / pdbx_entity_instance_feature / pdbx_entity_nonpoly / pdbx_nonpoly_scheme / pdbx_struct_assembly_gen / pdbx_struct_special_symmetry / pdbx_validate_rmsd_angle / refine / refine_hist / refine_ls_shell / reflns / reflns_shell / struct_asym / struct_conn Item _atom_site.B_iso_or_equiv / _atom_site.Cartn_x ... _atom_site.B_iso_or_equiv / _atom_site.Cartn_x / _atom_site.Cartn_y / _atom_site.Cartn_z / _atom_site.auth_atom_id / _atom_site.auth_comp_id / _atom_site.auth_seq_id / _atom_site.group_PDB / _atom_site.label_alt_id / _atom_site.label_asym_id / _atom_site.label_atom_id / _atom_site.label_comp_id / _atom_site.label_entity_id / _atom_site.label_seq_id / _atom_site.occupancy / _atom_site.type_symbol / _chem_comp.formula / _chem_comp.formula_weight / _chem_comp.id / _chem_comp.mon_nstd_flag / _chem_comp.name / _chem_comp.pdbx_synonyms / _chem_comp.type / _entity_src_gen.gene_src_common_name / _pdbx_struct_assembly_gen.asym_id_list / _pdbx_struct_special_symmetry.label_asym_id / _refine.B_iso_max / _refine.B_iso_mean / _refine.B_iso_min / _refine.ls_R_factor_R_free / _refine.ls_R_factor_R_work / _refine.ls_R_factor_obs / _refine.ls_number_reflns_R_work / _refine.ls_percent_reflns_obs / _refine_hist.cycle_id / _refine_hist.number_atoms_total / _refine_hist.pdbx_B_iso_mean_ligand / _refine_hist.pdbx_B_iso_mean_solvent / _refine_hist.pdbx_number_atoms_ligand / _refine_hist.pdbx_number_residues_total / _refine_ls_shell.R_factor_R_free_error / _refine_ls_shell.number_reflns_all / _refine_ls_shell.pdbx_total_number_of_bins_used / _reflns.pdbx_CC_half / _reflns.pdbx_Rrim_I_all / _reflns.pdbx_chi_squared / _reflns.pdbx_number_measured_all / _reflns.pdbx_redundancy / _reflns.pdbx_scaling_rejects Revision 3.1 Nov 29, 2023 Group / Refinement descriptionCategory / chem_comp_bond / pdbx_initial_refinement_modelRevision 3.2 Nov 6, 2024 Group / Category / pdbx_modification_feature / Item
Show all Show less
 Yorodumi
Yorodumi Open data
Open data Basic information
Basic information Components
Components Keywords
Keywords Function and homology information
Function and homology information
 X-RAY DIFFRACTION /
X-RAY DIFFRACTION /  SYNCHROTRON /
SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.59 Å
MOLECULAR REPLACEMENT / Resolution: 1.59 Å  Authors
Authors China, 1items
China, 1items  Citation
Citation Journal: Protein Cell / Year: 2022
Journal: Protein Cell / Year: 2022 Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 7vh8.cif.gz
7vh8.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb7vh8.ent.gz
pdb7vh8.ent.gz PDB format
PDB format 7vh8.json.gz
7vh8.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads https://data.pdbj.org/pub/pdb/validation_reports/vh/7vh8
https://data.pdbj.org/pub/pdb/validation_reports/vh/7vh8 ftp://data.pdbj.org/pub/pdb/validation_reports/vh/7vh8
ftp://data.pdbj.org/pub/pdb/validation_reports/vh/7vh8
 Links
Links Assembly
Assembly

 Components
Components

 X-RAY DIFFRACTION / Number of used crystals: 1
X-RAY DIFFRACTION / Number of used crystals: 1  Sample preparation
Sample preparation SYNCHROTRON / Site:
SYNCHROTRON / Site:  SSRF
SSRF  / Beamline: BL19U1 / Wavelength: 0.97852 Å
/ Beamline: BL19U1 / Wavelength: 0.97852 Å Processing
Processing MOLECULAR REPLACEMENT
MOLECULAR REPLACEMENT Movie
Movie Controller
Controller
















 PDBj
PDBj