+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 7pxw | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
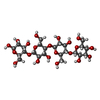
| Title | LPMO, expressed in E.coli, in complex with Cellotetraose | |||||||||
 Components Components | Auxiliary activity 9 | |||||||||
 Keywords Keywords | OXIDOREDUCTASE / Copper binding protein / Beta-sandwich fold / Auxillary | |||||||||
| Function / homology |  Function and homology information Function and homology informationlytic cellulose monooxygenase (C4-dehydrogenating) / cellulose catabolic process / monooxygenase activity / extracellular region / metal ion binding Similarity search - Function | |||||||||
| Biological species |  Lentinus similis (fungus) Lentinus similis (fungus) | |||||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  FOURIER SYNTHESIS / Resolution: 1.4 Å FOURIER SYNTHESIS / Resolution: 1.4 Å | |||||||||
 Authors Authors | Banerjee, S. / Muderspach, S.J. / Tandrup, T. / Ipsen, J. / Rollan, C.H. / Norholm, M. / Johansen, K.S. / Lo Leggio, L. | |||||||||
| Funding support |  Denmark, 2items Denmark, 2items
| |||||||||
 Citation Citation |  Journal: Iucrj / Year: 2022 Journal: Iucrj / Year: 2022Title: Changes in active-site geometry on X-ray photoreduction of a lytic polysaccharide monooxygenase active-site copper and saccharide binding. Authors: Tandrup, T. / Muderspach, S.J. / Banerjee, S. / Santoni, G. / Ipsen, J.O. / Hernandez-Rollan, C. / Norholm, M.H.H. / Johansen, K.S. / Meilleur, F. / Lo Leggio, L. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  7pxw.cif.gz 7pxw.cif.gz | 123.4 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb7pxw.ent.gz pdb7pxw.ent.gz | 93.5 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  7pxw.json.gz 7pxw.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/px/7pxw https://data.pdbj.org/pub/pdb/validation_reports/px/7pxw ftp://data.pdbj.org/pub/pdb/validation_reports/px/7pxw ftp://data.pdbj.org/pub/pdb/validation_reports/px/7pxw | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  7pqrC  7pxiC  7pxjC  7pxkC  7pxlC  7pxmC  7pxnC  7pxrC  7pxsC  7pxtC  7pxuC  7pxvC  7pydC  7pyeC  7pyfC  7pygC  7pyhC  7pyiC  7pylC  7pymC  7pynC  7pyoC  7pypC  7pyqC  7pyuC  7pywC  7pyxC  7pyyC  7pyzC  7pz0C  7pz3C  7pz4C  7pz5C  7pz6C  7pz7C  7pz8C  5achS C: citing same article ( S: Starting model for refinement |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
-Protein / Sugars , 2 types, 2 molecules A
| #1: Protein | Mass: 25259.832 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Lentinus similis (fungus) Lentinus similis (fungus)Production host:  References: UniProt: A0A0S2GKZ1 |
|---|---|
| #2: Polysaccharide | beta-D-glucopyranose-(1-4)-beta-D-glucopyranose-(1-4)-beta-D-glucopyranose-(1-4)-beta-D-glucopyranose |
-Non-polymers , 4 types, 324 molecules 






| #3: Chemical | | #4: Chemical | ChemComp-CU / | #5: Chemical | ChemComp-CL / #6: Water | ChemComp-HOH / | |
|---|
-Details
| Has ligand of interest | Y |
|---|---|
| Has protein modification | Y |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.57 Å3/Da / Density % sol: 52.2 % |
|---|---|
| Crystal grow | Temperature: 277 K / Method: batch mode / pH: 4.2 Details: 2.3 M ammonium sulfate, 0.1 M sodium acetate pH 4.2. Crystals were soaked in 0.05 M cellotetraose in crystallization conditions with 18% glycerol for 30 min. |
-Data collection
| Diffraction | Mean temperature: 100 K / Serial crystal experiment: N |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  MAX IV MAX IV  / Beamline: BioMAX / Wavelength: 0.98 Å / Beamline: BioMAX / Wavelength: 0.98 Å |
| Detector | Type: DECTRIS EIGER X 16M / Detector: PIXEL / Date: Dec 17, 2020 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.98 Å / Relative weight: 1 |
| Reflection | Resolution: 1.4→44.58 Å / Num. obs: 49868 / % possible obs: 99.6 % / Redundancy: 13.62 % / CC1/2: 0.999 / Rrim(I) all: 0.117 / Net I/σ(I): 12.07 |
| Reflection shell | Resolution: 1.4→1.44 Å / Redundancy: 13.57 % / Num. unique obs: 3724 / CC1/2: 0.55 / Rrim(I) all: 3.417 / % possible all: 99.7 |
- Processing
Processing
| Software |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  FOURIER SYNTHESIS FOURIER SYNTHESISStarting model: 5ACH Resolution: 1.4→44.58 Å / Cor.coef. Fo:Fc: 0.987 / Cor.coef. Fo:Fc free: 0.979 / SU B: 2.54 / SU ML: 0.042 / Cross valid method: THROUGHOUT / σ(F): 0 / ESU R: 0.047 / ESU R Free: 0.049 / Stereochemistry target values: MAXIMUM LIKELIHOOD Details: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS U VALUES : REFINED INDIVIDUALLY
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Ion probe radii: 0.8 Å / Shrinkage radii: 0.8 Å / VDW probe radii: 1.2 Å / Solvent model: MASK | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso max: 99.43 Å2 / Biso mean: 21.144 Å2 / Biso min: 13.71 Å2
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: final / Resolution: 1.4→44.58 Å
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 1.4→1.436 Å / Rfactor Rfree error: 0 / Total num. of bins used: 20
|
 Movie
Movie Controller
Controller



 PDBj
PDBj