+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 7kg8 | ||||||
|---|---|---|---|---|---|---|---|
| Title | Structure of human PARG complexed with PARG-061 | ||||||
 Components Components | Poly(ADP-ribose) glycohydrolase | ||||||
 Keywords Keywords | HYDROLASE/HYDROLASE inhibitor / HYDROLASE / hydrolase inhibitor complex / methyl xanthine / HYDROLASE-HYDROLASE inhibitor complex | ||||||
| Function / homology |  Function and homology information Function and homology informationnucleotide-sugar metabolic process / poly(ADP-ribose) glycohydrolase / poly(ADP-ribose) glycohydrolase activity / ATP generation from poly-ADP-D-ribose / POLB-Dependent Long Patch Base Excision Repair / regulation of DNA repair / base-excision repair, gap-filling / carbohydrate metabolic process / nuclear body / mitochondrial matrix ...nucleotide-sugar metabolic process / poly(ADP-ribose) glycohydrolase / poly(ADP-ribose) glycohydrolase activity / ATP generation from poly-ADP-D-ribose / POLB-Dependent Long Patch Base Excision Repair / regulation of DNA repair / base-excision repair, gap-filling / carbohydrate metabolic process / nuclear body / mitochondrial matrix / nucleoplasm / nucleus / cytoplasm / cytosol Similarity search - Function | ||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.43 Å MOLECULAR REPLACEMENT / Resolution: 1.43 Å | ||||||
 Authors Authors | Brosey, C.A. / Balapiti-Modarage, L.P.F. / Warden, L.S. / Jones, D.E. / Ahmed, Z. / Tainer, J.A. | ||||||
| Funding support |  United States, 1items United States, 1items
| ||||||
 Citation Citation |  Journal: Prog.Biophys.Mol.Biol. / Year: 2021 Journal: Prog.Biophys.Mol.Biol. / Year: 2021Title: Targeting SARS-CoV-2 Nsp3 macrodomain structure with insights from human poly(ADP-ribose) glycohydrolase (PARG) structures with inhibitors. Authors: Brosey, C.A. / Houl, J.H. / Katsonis, P. / Balapiti-Modarage, L.P.F. / Bommagani, S. / Arvai, A. / Moiani, D. / Bacolla, A. / Link, T. / Warden, L.S. / Lichtarge, O. / Jones, D.E. / Ahmed, Z. / Tainer, J.A. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  7kg8.cif.gz 7kg8.cif.gz | 381.1 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb7kg8.ent.gz pdb7kg8.ent.gz | 256.9 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  7kg8.json.gz 7kg8.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/kg/7kg8 https://data.pdbj.org/pub/pdb/validation_reports/kg/7kg8 ftp://data.pdbj.org/pub/pdb/validation_reports/kg/7kg8 ftp://data.pdbj.org/pub/pdb/validation_reports/kg/7kg8 | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  7kfpC  7kg0C  7kg1C  7kg3C  7kg6C  7kg7C  7kxbC  7lg7C  4b1gS C: citing same article ( S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||||||
| Unit cell |
|
- Components
Components
-Protein , 1 types, 1 molecules A
| #1: Protein | Mass: 61096.492 Da / Num. of mol.: 1 / Mutation: K616A, Q617A, K618A, E688A, K689A, K690A Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Gene: PARG / Production host: Homo sapiens (human) / Gene: PARG / Production host:  References: UniProt: Q86W56, poly(ADP-ribose) glycohydrolase |
|---|
-Non-polymers , 5 types, 442 molecules 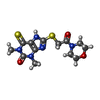








| #2: Chemical | ChemComp-WDM / | ||||||
|---|---|---|---|---|---|---|---|
| #3: Chemical | ChemComp-DMS / #4: Chemical | #5: Chemical | ChemComp-CAC / | #6: Water | ChemComp-HOH / | |
-Details
| Has ligand of interest | Y |
|---|---|
| Has protein modification | Y |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.33 Å3/Da / Density % sol: 47.2 % |
|---|---|
| Crystal grow | Temperature: 295 K / Method: vapor diffusion, hanging drop / pH: 7.5 / Details: 0.1 M PCTP, pH 7.5, 0.2 M AmSO4, 18-23% PEG3350 |
-Data collection
| Diffraction | Mean temperature: 100 K / Serial crystal experiment: N |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  SSRL SSRL  / Beamline: BL9-2 / Wavelength: 0.9795 Å / Beamline: BL9-2 / Wavelength: 0.9795 Å |
| Detector | Type: DECTRIS PILATUS 6M / Detector: PIXEL / Date: Jan 17, 2019 |
| Radiation | Monochromator: Si(111) and Si(220) double crystal / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.9795 Å / Relative weight: 1 |
| Reflection | Resolution: 1.43→38.33 Å / Num. obs: 103197 / % possible obs: 99.58 % / Redundancy: 7.8 % / Biso Wilson estimate: 22.04 Å2 / CC1/2: 0.999 / CC star: 1 / Rmerge(I) obs: 0.06384 / Rpim(I) all: 0.02406 / Rrim(I) all: 0.06836 / Net I/σ(I): 13.49 |
| Reflection shell | Resolution: 1.43→1.481 Å / Redundancy: 8.1 % / Rmerge(I) obs: 2.646 / Mean I/σ(I) obs: 0.81 / Num. unique obs: 10175 / CC1/2: 0.393 / CC star: 0.751 / Rpim(I) all: 0.9804 / Rrim(I) all: 2.826 / % possible all: 98.92 |
- Processing
Processing
| Software |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 4B1G Resolution: 1.43→38.33 Å / SU ML: 0.1635 / Cross valid method: FREE R-VALUE / σ(F): 1.34 / Phase error: 20.9692 Stereochemistry target values: GeoStd + Monomer Library + CDL v1.2
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Shrinkage radii: 0.9 Å / VDW probe radii: 1.11 Å / Solvent model: FLAT BULK SOLVENT MODEL | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 28.88 Å2 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.43→38.33 Å
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS params. | Method: refined / Origin x: 17.7408166808 Å / Origin y: -2.71161868583 Å / Origin z: -8.53140189802 Å
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS group | Selection details: all |
 Movie
Movie Controller
Controller













 PDBj
PDBj









