[English] 日本語
 Yorodumi
Yorodumi- PDB-7d37: Solution structure of Acm2-precursor peptide of Heat-stable enter... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 7d37 | ||||||
|---|---|---|---|---|---|---|---|
| Title | Solution structure of Acm2-precursor peptide of Heat-stable enterotoxin produced by Enterotoxigenic Escherichia coli | ||||||
 Components Components | CYS-CY1-GLU-LEU-CYS-CYS-ASN-PRO-ALA-CY1-THR-GLY-CYS | ||||||
 Keywords Keywords | TOXIN / Heat-stable enterotoxin / topological isomer / STh / STh(6-18) | ||||||
| Function / homology | Heat-stable enterotoxin, STa / Heat-stable enterotoxin, conserved site / Heat-stable enterotoxin ST / Heat-stable enterotoxins signature. / toxin activity / extracellular space / Heat-stable enterotoxin A3/A4 Function and homology information Function and homology information | ||||||
| Biological species |  | ||||||
| Method | SOLUTION NMR / simulated annealing | ||||||
 Authors Authors | Shimamoto, S. / Hidaka, Y. | ||||||
 Citation Citation |  Journal: Molecules / Year: 2020 Journal: Molecules / Year: 2020Title: Topological Regulation of the Bioactive Conformation of a Disulfide-Rich Peptide, Heat-Stable Enterotoxin. Authors: Shimamoto, S. / Fukutsuji, M. / Osumi, T. / Goto, M. / Toyoda, H. / Hidaka, Y. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  7d37.cif.gz 7d37.cif.gz | 37 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb7d37.ent.gz pdb7d37.ent.gz | 24.3 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  7d37.json.gz 7d37.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Summary document |  7d37_validation.pdf.gz 7d37_validation.pdf.gz | 368.4 KB | Display |  wwPDB validaton report wwPDB validaton report |
|---|---|---|---|---|
| Full document |  7d37_full_validation.pdf.gz 7d37_full_validation.pdf.gz | 404.4 KB | Display | |
| Data in XML |  7d37_validation.xml.gz 7d37_validation.xml.gz | 5.2 KB | Display | |
| Data in CIF |  7d37_validation.cif.gz 7d37_validation.cif.gz | 6.9 KB | Display | |
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/d3/7d37 https://data.pdbj.org/pub/pdb/validation_reports/d3/7d37 ftp://data.pdbj.org/pub/pdb/validation_reports/d3/7d37 ftp://data.pdbj.org/pub/pdb/validation_reports/d3/7d37 | HTTPS FTP |
-Related structure data
| Related structure data |  7cssC C: citing same article ( |
|---|---|
| Similar structure data | |
| Other databases |
|
- Links
Links
- Assembly
Assembly
| Deposited unit | 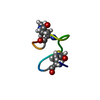
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein/peptide | Mass: 1461.751 Da / Num. of mol.: 1 / Source method: obtained synthetically / Source: (synth.)  |
|---|---|
| Has ligand of interest | Y |
| Has protein modification | Y |
-Experimental details
-Experiment
| Experiment | Method: SOLUTION NMR | ||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
|
- Sample preparation
Sample preparation
| Details |
| ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sample |
| ||||||||||||||||||
| Sample conditions | Ionic strength: 1 mM / Label: conditions_1 / pH: 3 / Pressure: 1.0 atm / Temperature: 298.15 K |
-NMR measurement
| NMR spectrometer | Type: JEOL JNM ECA800 / Manufacturer: JEOL / Model: JNM ECA800 / Field strength: 800 MHz |
|---|
- Processing
Processing
| NMR software | Name: CNS / Developer: Brunger, Adams, Clore, Gros, Nilges and Read / Classification: structure calculation |
|---|---|
| Refinement | Method: simulated annealing / Software ordinal: 4 |
| NMR representative | Selection criteria: lowest energy |
| NMR ensemble | Conformer selection criteria: structures with the lowest energy Conformers calculated total number: 1000 / Conformers submitted total number: 10 |
 Movie
Movie Controller
Controller








 PDBj
PDBj