+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 7bvw | ||||||
|---|---|---|---|---|---|---|---|
| Title | Crystal structure of the RING-H2 domain of Arabidopsis RMR1 | ||||||
 Components Components | AT5G66160 protein | ||||||
 Keywords Keywords | TRANSPORT PROTEIN / C3H2C3 / ring finger / E3 ligase | ||||||
| Function / homology | : / Ring finger domain / Ring finger / Zinc finger RING-type profile. / Zinc finger, RING-type / Zinc finger, RING/FYVE/PHD-type / membrane / AT5G66160 protein Function and homology information Function and homology information | ||||||
| Biological species |  | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  MOLECULAR REPLACEMENT / Resolution: 2.1 Å MOLECULAR REPLACEMENT / Resolution: 2.1 Å | ||||||
 Authors Authors | Chen, S. / Wong, K.B. | ||||||
 Citation Citation |  Journal: J.Biol.Chem. / Year: 2025 Journal: J.Biol.Chem. / Year: 2025Title: The RING-finger domain of Arabidopsis RMR functions as an E3 ligase essential for post-Golgi trafficking. Authors: Chen, S. / Zeng, Y.L. / Wong, H.Y. / Chen, Y.H. / Yang, L. / Luo, F. / Gao, C.J. / Jiang, L.W. / Wong, K.B. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  7bvw.cif.gz 7bvw.cif.gz | 78.6 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb7bvw.ent.gz pdb7bvw.ent.gz | 57.7 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  7bvw.json.gz 7bvw.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/bv/7bvw https://data.pdbj.org/pub/pdb/validation_reports/bv/7bvw ftp://data.pdbj.org/pub/pdb/validation_reports/bv/7bvw ftp://data.pdbj.org/pub/pdb/validation_reports/bv/7bvw | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  4v3kS S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 | 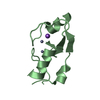
| ||||||||
| Unit cell |
| ||||||||
| Components on special symmetry positions |
|
- Components
Components
| #1: Protein | Mass: 9220.559 Da / Num. of mol.: 2 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   #2: Chemical | ChemComp-ZN / #3: Chemical | ChemComp-NA / | #4: Water | ChemComp-HOH / | Has ligand of interest | Y | Has protein modification | N | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 1.8 Å3/Da / Density % sol: 31.67 % |
|---|---|
| Crystal grow | Temperature: 289 K / Method: vapor diffusion, hanging drop / pH: 5.5 Details: 0.2 M Ammonium acetate, 0.1 M BIS-TRIS pH 5.5, 20% w/v Polyethylene glycol 3,350 |
-Data collection
| Diffraction | Mean temperature: 93 K / Serial crystal experiment: N |
|---|---|
| Diffraction source | Source:  ROTATING ANODE / Type: RIGAKU FR-E+ SUPERBRIGHT / Wavelength: 1.54 Å ROTATING ANODE / Type: RIGAKU FR-E+ SUPERBRIGHT / Wavelength: 1.54 Å |
| Detector | Type: RIGAKU RAXIS IV++ / Detector: IMAGE PLATE / Date: Jan 4, 2018 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.54 Å / Relative weight: 1 |
| Reflection | Resolution: 2.1→37.72 Å / Num. obs: 7849 / % possible obs: 98.5 % / Redundancy: 3.6 % / CC1/2: 0.992 / Rmerge(I) obs: 0.098 / Rpim(I) all: 0.061 / Rrim(I) all: 0.116 / Net I/σ(I): 9 / Num. measured all: 27884 |
| Reflection shell | Resolution: 2.1→2.16 Å / Redundancy: 3.4 % / Rmerge(I) obs: 0.213 / Num. unique obs: 650 / CC1/2: 0.95 / Rpim(I) all: 0.135 / Rrim(I) all: 0.253 / % possible all: 97.4 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 4V3K Resolution: 2.1→32.23 Å / SU ML: 0.22 / Cross valid method: THROUGHOUT / σ(F): 1.36 / Phase error: 25.31
| ||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Shrinkage radii: 0.9 Å / VDW probe radii: 1.11 Å | ||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso max: 87.05 Å2 / Biso mean: 27.392 Å2 / Biso min: 12.33 Å2 | ||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: final / Resolution: 2.1→32.23 Å
| ||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Refine-ID: X-RAY DIFFRACTION / Rfactor Rfree error: 0
| ||||||||||||||||||||||||||||||||||||||||
| Refinement TLS params. | Method: refined / Origin x: 1.4047 Å / Origin y: 5.8871 Å / Origin z: 94.4639 Å
| ||||||||||||||||||||||||||||||||||||||||
| Refinement TLS group |
|
 Movie
Movie Controller
Controller













 PDBj
PDBj






