[English] 日本語
 Yorodumi
Yorodumi- PDB-6rha: Crystal structure of the amyloid-like NTVTFN segment from the Can... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 6rha | ||||||
|---|---|---|---|---|---|---|---|
| Title | Crystal structure of the amyloid-like NTVTFN segment from the Candida albicans Agglutinin-like protein (Adhesin) 5 | ||||||
 Components Components | Agglutinin-like protein 5 | ||||||
 Keywords Keywords | PROTEIN FIBRIL / Bacterial steric-zipper cross-beta amyloid fibril from Candida albicans | ||||||
| Function / homology |  Function and homology information Function and homology informationcell adhesion involved in multi-species biofilm formation / hyphal growth / single-species biofilm formation on inanimate substrate / yeast-form cell wall / hyphal cell wall / cell adhesion involved in single-species biofilm formation / side of membrane / cell-cell adhesion / extracellular vesicle / cell surface / plasma membrane Similarity search - Function | ||||||
| Biological species |  Candida albicans (yeast) Candida albicans (yeast) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / MOLECULAR REPLACEMENT /  molecular replacement / Resolution: 1.6 Å molecular replacement / Resolution: 1.6 Å | ||||||
| Model details | Adhesin | ||||||
 Authors Authors | Landau, M. / Perov, S. | ||||||
 Citation Citation |  Journal: To be published Journal: To be publishedTitle: Amyloid structures from a Candida albicans adhesin Structure and conservation of amyloid spines from fungal adhesins Authors: Perov, S. / Landau, M. / Lipke, P.N. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  6rha.cif.gz 6rha.cif.gz | 11.4 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb6rha.ent.gz pdb6rha.ent.gz | 6 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  6rha.json.gz 6rha.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/rh/6rha https://data.pdbj.org/pub/pdb/validation_reports/rh/6rha ftp://data.pdbj.org/pub/pdb/validation_reports/rh/6rha ftp://data.pdbj.org/pub/pdb/validation_reports/rh/6rha | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
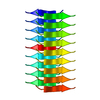
| 1 | x 9
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein/peptide | Mass: 694.733 Da / Num. of mol.: 2 Fragment: Amyloid spine segment NTVTFN from Als5 (residues 156-161) secreted by Candida albicans Source method: obtained synthetically / Details: NTVTFN from Als5, synthesized / Source: (synth.)  Candida albicans (yeast) / References: UniProt: O13368 Candida albicans (yeast) / References: UniProt: O13368#2: Water | ChemComp-HOH / | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal grow | Temperature: 293 K / Method: vapor diffusion, hanging drop / pH: 7.5 Details: Reservoir contained 0.1M HEPES 7.5 pH, 0.8 M NaH2PO4, and 0.8M KH2PO4 |
|---|
-Data collection
| Diffraction | Mean temperature: 100 K / Serial crystal experiment: N | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  ESRF ESRF  / Beamline: ID23-2 / Wavelength: 0.8729 Å / Beamline: ID23-2 / Wavelength: 0.8729 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Detector | Type: DECTRIS PILATUS3 2M / Detector: PIXEL / Date: May 10, 2015 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Radiation wavelength | Wavelength: 0.8729 Å / Relative weight: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reflection | Resolution: 1.6→19.91 Å / Num. obs: 991 / % possible obs: 98.2 % / Redundancy: 14.28 % / Biso Wilson estimate: 19.469 Å2 / CC1/2: 0.975 / Rmerge(I) obs: 0.27 / Rrim(I) all: 0.28 / Χ2: 0.747 / Net I/σ(I): 7.3 / Num. measured all: 14151 / Scaling rejects: 28 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reflection shell | Diffraction-ID: 1
|
-Phasing
| Phasing | Method:  molecular replacement molecular replacement | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Phasing MR | Model details: Phaser MODE: MR_AUTO / Packing: 0
|
- Processing
Processing
| Software |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: beta strand Resolution: 1.6→19.91 Å / Cor.coef. Fo:Fc: 0.939 / Cor.coef. Fo:Fc free: 0.928 / SU B: 2.893 / SU ML: 0.093 / SU R Cruickshank DPI: 0.1536 / Cross valid method: THROUGHOUT / σ(F): 0 / ESU R: 0.154 / ESU R Free: 0.121 Details: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS U VALUES : REFINED INDIVIDUALLY
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Ion probe radii: 0.8 Å / Shrinkage radii: 0.8 Å / VDW probe radii: 1.2 Å | |||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso max: 31.18 Å2 / Biso mean: 10.562 Å2 / Biso min: 6.63 Å2
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: final / Resolution: 1.6→19.91 Å
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 1.601→1.643 Å / Rfactor Rfree error: 0 / Total num. of bins used: 20
|
 Movie
Movie Controller
Controller











 PDBj
PDBj
