+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 6cct | ||||||
|---|---|---|---|---|---|---|---|
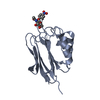
| Title | Fragment of GID4 in complex with a short peptide | ||||||
 Components Components |
| ||||||
 Keywords Keywords | PEPTIDE BINDING PROTEIN / Structural Genomics / Structural Genomics Consortium / SGC | ||||||
| Function / homology | Vacuolar import/degradation protein Vid24 / Vacuolar import and degradation protein / ubiquitin ligase complex / Regulation of pyruvate metabolism / ubiquitin protein ligase activity / proteasome-mediated ubiquitin-dependent protein catabolic process / cytosol / Glucose-induced degradation protein 4 homolog Function and homology information Function and homology information | ||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human)synthetic construct (others) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2.4 Å MOLECULAR REPLACEMENT / Resolution: 2.4 Å | ||||||
 Authors Authors | Dong, C. / Tempel, W. / Bountra, C. / Arrowsmith, C.H. / Edwards, A.M. / Min, J. / Structural Genomics Consortium (SGC) | ||||||
 Citation Citation |  Journal: Nat. Chem. Biol. / Year: 2018 Journal: Nat. Chem. Biol. / Year: 2018Title: Molecular basis of GID4-mediated recognition of degrons for the Pro/N-end rule pathway. Authors: Dong, C. / Zhang, H. / Li, L. / Tempel, W. / Loppnau, P. / Min, J. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  6cct.cif.gz 6cct.cif.gz | 79.7 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb6cct.ent.gz pdb6cct.ent.gz | 58.7 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  6cct.json.gz 6cct.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/cc/6cct https://data.pdbj.org/pub/pdb/validation_reports/cc/6cct ftp://data.pdbj.org/pub/pdb/validation_reports/cc/6cct ftp://data.pdbj.org/pub/pdb/validation_reports/cc/6cct | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  6ccrSC  6ccuC  6cd8C  6cd9C  6cdcC  6cdgC S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 19604.777 Da / Num. of mol.: 1 / Fragment: residues 124-289 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Gene: GID4, C17orf39, VID24 / Production host: Homo sapiens (human) / Gene: GID4, C17orf39, VID24 / Production host:  |
|---|---|
| #2: Protein/peptide | Mass: 428.522 Da / Num. of mol.: 1 / Source method: obtained synthetically / Source: (synth.) synthetic construct (others) |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.05 Å3/Da / Density % sol: 40 % / Mosaicity: 0.23 ° |
|---|---|
| Crystal grow | Temperature: 291 K / Method: vapor diffusion, sitting drop Details: 20% PEG3350, 2% Tacsimate pH 7.0 and 0.1M HEPES pH 7.5 |
-Data collection
| Diffraction | Mean temperature: 100 K | |||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  APS APS  / Beamline: 24-ID-E / Wavelength: 0.97918 Å / Beamline: 24-ID-E / Wavelength: 0.97918 Å | |||||||||||||||||||||||||||||||||
| Detector | Type: DECTRIS EIGER X 16M / Detector: PIXEL / Date: Jun 12, 2017 | |||||||||||||||||||||||||||||||||
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray | |||||||||||||||||||||||||||||||||
| Radiation wavelength | Wavelength: 0.97918 Å / Relative weight: 1 | |||||||||||||||||||||||||||||||||
| Reflection | Resolution: 2.1→101.67 Å / Num. obs: 10614 / % possible obs: 100 % / Redundancy: 11.8 % / CC1/2: 0.999 / Rmerge(I) obs: 0.051 / Rpim(I) all: 0.016 / Rrim(I) all: 0.053 / Net I/σ(I): 22.4 / Num. measured all: 125391 / Scaling rejects: 0 | |||||||||||||||||||||||||||||||||
| Reflection shell | Diffraction-ID: 1
|
- Processing
Processing
| Software |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: early version of model from PDB entry 6CCR Resolution: 2.4→39.3 Å / Cor.coef. Fo:Fc: 0.942 / Cor.coef. Fo:Fc free: 0.935 / SU B: 18.466 / SU ML: 0.196 / Cross valid method: THROUGHOUT / σ(F): 0 / ESU R: 0.52 / ESU R Free: 0.286 / Stereochemistry target values: MAXIMUM LIKELIHOOD Details: The rotamer of peptide residue T2 was not clearly resolved by electron density, but chosen so as to enable plausible interactions with surrounding atoms of the model.
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Ion probe radii: 0.8 Å / Shrinkage radii: 0.8 Å / VDW probe radii: 1.2 Å / Solvent model: MASK | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso max: 134.05 Å2 / Biso mean: 60.158 Å2 / Biso min: 38.87 Å2
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: final / Resolution: 2.4→39.3 Å
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 2.4→2.462 Å / Total num. of bins used: 20
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS params. | Method: refined / Refine-ID: X-RAY DIFFRACTION
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS group |
|
 Movie
Movie Controller
Controller











 PDBj
PDBj