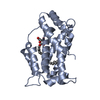
Entry Database : PDB / ID : 6b8lTitle Crystal Structure of the Apo/CaM:Kv7.4 (KCNQ4) AB Domain Complex Calmodulin-1 Potassium voltage-gated channel subfamily KQT member 4 Keywords / / Function / homology Function Domain/homology Component
/ / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / Biological species Homo sapiens (human)Method / / Resolution : 2.3 Å Authors Chang, A. / Abderemane-Ali, F. / Minor, D.L. Funding support Organization Grant number Country National Institutes of Health/National Institute on Deafness and Other Communication Disorders (NIH/NIDCD) DC007664
Journal : Neuron / Year : 2018Title : A Calmodulin C-Lobe Ca2+-Dependent Switch Governs Kv7 Channel FunctionAuthors : Chang, A. / Abderemane-Ali, F. / Hura, G.L. / Rossen, N.D. / Gate, R.E. / Minor, D.L. History Deposition Oct 9, 2017 Deposition site / Processing site Revision 1.0 Mar 14, 2018 Provider / Type Revision 1.1 Dec 18, 2019 Group / Category Item / _pdbx_audit_support.grant_numberRevision 1.2 Mar 13, 2024 Group / Data collection / Database referencesCategory chem_comp_atom / chem_comp_bond ... chem_comp_atom / chem_comp_bond / database_2 / pdbx_unobs_or_zero_occ_atoms Item / _database_2.pdbx_database_accession
Show all Show less
 Open data
Open data Basic information
Basic information Components
Components Keywords
Keywords Function and homology information
Function and homology information Homo sapiens (human)
Homo sapiens (human) X-RAY DIFFRACTION /
X-RAY DIFFRACTION /  SYNCHROTRON / Resolution: 2.3 Å
SYNCHROTRON / Resolution: 2.3 Å  Authors
Authors United States, 1items
United States, 1items  Citation
Citation Journal: Neuron / Year: 2018
Journal: Neuron / Year: 2018 Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 6b8l.cif.gz
6b8l.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb6b8l.ent.gz
pdb6b8l.ent.gz PDB format
PDB format 6b8l.json.gz
6b8l.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads https://data.pdbj.org/pub/pdb/validation_reports/b8/6b8l
https://data.pdbj.org/pub/pdb/validation_reports/b8/6b8l ftp://data.pdbj.org/pub/pdb/validation_reports/b8/6b8l
ftp://data.pdbj.org/pub/pdb/validation_reports/b8/6b8l Links
Links Assembly
Assembly




 Components
Components Homo sapiens (human) / Gene: KCNQ4 / Plasmid: pET28 / Production host:
Homo sapiens (human) / Gene: KCNQ4 / Plasmid: pET28 / Production host: 
 Homo sapiens (human) / Gene: CALM1, CALM, CAM, CAM1 / Plasmid: pEGST / Production host:
Homo sapiens (human) / Gene: CALM1, CALM, CAM, CAM1 / Plasmid: pEGST / Production host: 
 X-RAY DIFFRACTION / Number of used crystals: 1
X-RAY DIFFRACTION / Number of used crystals: 1  Sample preparation
Sample preparation Processing
Processing Movie
Movie Controller
Controller
















 PDBj
PDBj

























