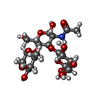
Entry Database : PDB / ID : 5zvcTitle P domain of GII.13 norovirus capsid complexed with Lewis A trisaccharide Major capsid protein VP1 Keywords / / / / Function / homology Function Domain/homology Component
/ / / / / / / / / / / / / / / / / / / / Biological species Method / / Resolution : 1.8 Å Authors Chen, Y. / Li, X. Funding support Organization Grant number Country National Natural Science Foundation of China 31670752
Journal : J. Virol. / Year : 2019Title : Structural Adaptations of Norovirus GII.17/13/21 Lineage through Two Distinct Evolutionary Paths.Authors : Qian, Y. / Song, M. / Jiang, X. / Xia, M. / Meller, J. / Tan, M. / Chen, Y. / Li, X. / Rao, Z. History Deposition May 10, 2018 Deposition site / Processing site Revision 1.0 Oct 3, 2018 Provider / Type Revision 1.1 Apr 17, 2019 Group / Database references / Category / citation_authorItem _citation.country / _citation.journal_abbrev ... _citation.country / _citation.journal_abbrev / _citation.journal_id_ASTM / _citation.journal_id_CSD / _citation.journal_id_ISSN / _citation.journal_volume / _citation.pdbx_database_id_DOI / _citation.pdbx_database_id_PubMed / _citation.title / _citation.year Revision 2.0 Jul 29, 2020 Group Atomic model / Data collection ... Atomic model / Data collection / Derived calculations / Structure summary Category atom_site / atom_site_anisotrop ... atom_site / atom_site_anisotrop / chem_comp / entity / pdbx_branch_scheme / pdbx_chem_comp_identifier / pdbx_entity_branch / pdbx_entity_branch_descriptor / pdbx_entity_branch_link / pdbx_entity_branch_list / pdbx_entity_nonpoly / pdbx_nonpoly_scheme / pdbx_struct_assembly_gen / struct_asym / struct_conn / struct_site / struct_site_gen Item _atom_site.B_iso_or_equiv / _atom_site.Cartn_x ... _atom_site.B_iso_or_equiv / _atom_site.Cartn_x / _atom_site.Cartn_y / _atom_site.Cartn_z / _atom_site.auth_asym_id / _atom_site.auth_atom_id / _atom_site.auth_comp_id / _atom_site.auth_seq_id / _atom_site.label_asym_id / _atom_site.label_atom_id / _atom_site.label_comp_id / _atom_site.label_entity_id / _atom_site.occupancy / _atom_site.type_symbol / _atom_site_anisotrop.U[1][1] / _atom_site_anisotrop.U[1][2] / _atom_site_anisotrop.U[1][3] / _atom_site_anisotrop.U[2][2] / _atom_site_anisotrop.U[2][3] / _atom_site_anisotrop.U[3][3] / _atom_site_anisotrop.pdbx_auth_asym_id / _atom_site_anisotrop.pdbx_auth_atom_id / _atom_site_anisotrop.pdbx_auth_comp_id / _atom_site_anisotrop.pdbx_auth_seq_id / _atom_site_anisotrop.pdbx_label_asym_id / _atom_site_anisotrop.pdbx_label_atom_id / _atom_site_anisotrop.pdbx_label_comp_id / _atom_site_anisotrop.type_symbol / _chem_comp.name / _chem_comp.type / _pdbx_struct_assembly_gen.asym_id_list / _struct_conn.pdbx_dist_value / _struct_conn.pdbx_leaving_atom_flag / _struct_conn.ptnr1_auth_asym_id / _struct_conn.ptnr1_auth_comp_id / _struct_conn.ptnr1_auth_seq_id / _struct_conn.ptnr1_label_asym_id / _struct_conn.ptnr1_label_atom_id / _struct_conn.ptnr1_label_comp_id / _struct_conn.ptnr2_auth_asym_id / _struct_conn.ptnr2_auth_comp_id / _struct_conn.ptnr2_auth_seq_id / _struct_conn.ptnr2_label_asym_id / _struct_conn.ptnr2_label_atom_id / _struct_conn.ptnr2_label_comp_id Description / Provider / Type Revision 3.0 Aug 5, 2020 Group Advisory / Atomic model ... Advisory / Atomic model / Data collection / Derived calculations / Structure summary Category atom_site / atom_site_anisotrop ... atom_site / atom_site_anisotrop / pdbx_distant_solvent_atoms / pdbx_molecule_features / pdbx_nonpoly_scheme / pdbx_validate_close_contact / struct_conn Item _atom_site.B_iso_or_equiv / _atom_site.Cartn_x ... _atom_site.B_iso_or_equiv / _atom_site.Cartn_x / _atom_site.Cartn_y / _atom_site.Cartn_z / _atom_site_anisotrop.U[1][1] / _atom_site_anisotrop.U[1][2] / _atom_site_anisotrop.U[1][3] / _atom_site_anisotrop.U[2][2] / _atom_site_anisotrop.U[2][3] / _atom_site_anisotrop.U[3][3] / _pdbx_distant_solvent_atoms.auth_seq_id / _pdbx_distant_solvent_atoms.neighbor_ligand_distance / _pdbx_distant_solvent_atoms.neighbor_macromolecule_distance / _pdbx_nonpoly_scheme.auth_seq_num / _pdbx_validate_close_contact.auth_seq_id_1 / _pdbx_validate_close_contact.auth_seq_id_2 / _struct_conn.pdbx_leaving_atom_flag Revision 3.1 Nov 22, 2023 Group Data collection / Database references ... Data collection / Database references / Refinement description / Structure summary Category chem_comp / chem_comp_atom ... chem_comp / chem_comp_atom / chem_comp_bond / database_2 / pdbx_initial_refinement_model / struct_ncs_dom_lim Item _chem_comp.pdbx_synonyms / _database_2.pdbx_DOI ... _chem_comp.pdbx_synonyms / _database_2.pdbx_DOI / _database_2.pdbx_database_accession / _struct_ncs_dom_lim.beg_auth_comp_id / _struct_ncs_dom_lim.beg_label_asym_id / _struct_ncs_dom_lim.beg_label_comp_id / _struct_ncs_dom_lim.beg_label_seq_id / _struct_ncs_dom_lim.end_auth_comp_id / _struct_ncs_dom_lim.end_label_asym_id / _struct_ncs_dom_lim.end_label_comp_id / _struct_ncs_dom_lim.end_label_seq_id
Show all Show less
 Yorodumi
Yorodumi Open data
Open data Basic information
Basic information Components
Components Keywords
Keywords Function and homology information
Function and homology information Norovirus Hu/GII.13/10N4598/2010/NP
Norovirus Hu/GII.13/10N4598/2010/NP X-RAY DIFFRACTION /
X-RAY DIFFRACTION /  MOLECULAR REPLACEMENT / Resolution: 1.8 Å
MOLECULAR REPLACEMENT / Resolution: 1.8 Å  Authors
Authors China, 1items
China, 1items  Citation
Citation Journal: J. Virol. / Year: 2019
Journal: J. Virol. / Year: 2019 Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 5zvc.cif.gz
5zvc.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb5zvc.ent.gz
pdb5zvc.ent.gz PDB format
PDB format 5zvc.json.gz
5zvc.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads https://data.pdbj.org/pub/pdb/validation_reports/zv/5zvc
https://data.pdbj.org/pub/pdb/validation_reports/zv/5zvc ftp://data.pdbj.org/pub/pdb/validation_reports/zv/5zvc
ftp://data.pdbj.org/pub/pdb/validation_reports/zv/5zvc




 Links
Links Assembly
Assembly
 Components
Components Norovirus Hu/GII.13/10N4598/2010/NP / Production host:
Norovirus Hu/GII.13/10N4598/2010/NP / Production host: 
 X-RAY DIFFRACTION / Number of used crystals: 1
X-RAY DIFFRACTION / Number of used crystals: 1  Sample preparation
Sample preparation ROTATING ANODE / Type: RIGAKU MICROMAX-007 / Wavelength: 1.5418 Å
ROTATING ANODE / Type: RIGAKU MICROMAX-007 / Wavelength: 1.5418 Å Processing
Processing MOLECULAR REPLACEMENT
MOLECULAR REPLACEMENT Movie
Movie Controller
Controller












 PDBj
PDBj