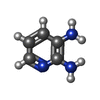
Entry Database : PDB / ID : 5ludTitle Structure of Cyclophilin A in complex with 2,3-Diaminopyridine Peptidyl-prolyl cis-trans isomerase Keywords / / / Function / homology Function Domain/homology Component
/ / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / Biological species Homo sapiens (human)Method / / / Resolution : 1.25 Å Authors McNae, I.W. / Nowicki, M.W. / Blackburn, E.A. / Wear, M.A. / Walkinshaw, M.D. Funding support Organization Grant number Country Wellcome Trust 081287/Z/06/Z Wellcome Trust 101527/Z/13/Z
Journal : FEBS Open Bio / Year : 2017Title : Thermo-kinetic analysis space expansion for cyclophilin-ligand interactions - identification of a new nonpeptide inhibitor using BiacoreTM T200.Authors : Wear, M.A. / Nowicki, M.W. / Blackburn, E.A. / McNae, I.W. / Walkinshaw, M.D. History Deposition Sep 8, 2016 Deposition site / Processing site Revision 1.0 Apr 5, 2017 Provider / Type Revision 1.1 Apr 19, 2017 Group Revision 1.2 Jan 17, 2024 Group / Database references / Refinement descriptionCategory chem_comp_atom / chem_comp_bond ... chem_comp_atom / chem_comp_bond / database_2 / pdbx_initial_refinement_model Item / _database_2.pdbx_database_accession
Show all Show less
 Open data
Open data Basic information
Basic information Components
Components Keywords
Keywords Function and homology information
Function and homology information Homo sapiens (human)
Homo sapiens (human) X-RAY DIFFRACTION /
X-RAY DIFFRACTION /  SYNCHROTRON /
SYNCHROTRON /  FOURIER SYNTHESIS / Resolution: 1.25 Å
FOURIER SYNTHESIS / Resolution: 1.25 Å  Authors
Authors United Kingdom, 2items
United Kingdom, 2items  Citation
Citation Journal: FEBS Open Bio / Year: 2017
Journal: FEBS Open Bio / Year: 2017 Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 5lud.cif.gz
5lud.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb5lud.ent.gz
pdb5lud.ent.gz PDB format
PDB format 5lud.json.gz
5lud.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads https://data.pdbj.org/pub/pdb/validation_reports/lu/5lud
https://data.pdbj.org/pub/pdb/validation_reports/lu/5lud ftp://data.pdbj.org/pub/pdb/validation_reports/lu/5lud
ftp://data.pdbj.org/pub/pdb/validation_reports/lu/5lud
 Links
Links Assembly
Assembly
 Components
Components Homo sapiens (human) / Gene: HEL-S-69p / Production host:
Homo sapiens (human) / Gene: HEL-S-69p / Production host: 
 X-RAY DIFFRACTION / Number of used crystals: 1
X-RAY DIFFRACTION / Number of used crystals: 1  Sample preparation
Sample preparation SYNCHROTRON / Site:
SYNCHROTRON / Site:  Diamond
Diamond  / Beamline: I02 / Wavelength: 0.9795 Å
/ Beamline: I02 / Wavelength: 0.9795 Å Processing
Processing FOURIER SYNTHESIS
FOURIER SYNTHESIS Movie
Movie Controller
Controller












 PDBj
PDBj