[English] 日本語
 Yorodumi
Yorodumi- PDB-5l7f: Crystal structure of MMP12 mutant K421A in complex with RXP470.1 ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 5l7f | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Crystal structure of MMP12 mutant K421A in complex with RXP470.1 conjugated with fluorophore Cy5,5 in space group P21. | |||||||||
 Components Components | Macrophage metalloelastase | |||||||||
 Keywords Keywords | HYDROLASE / Probes for optical imaging / MMP-12 / metalloprotease / inflammation / aneurysm / fluorophore / Cy5 / 5 / cyanine | |||||||||
| Function / homology |  Function and homology information Function and homology informationmacrophage elastase / negative regulation of endothelial cell-matrix adhesion / bronchiole development / positive regulation of epithelial cell proliferation involved in wound healing / elastin catabolic process / regulation of defense response to virus by host / positive regulation of type I interferon-mediated signaling pathway / wound healing, spreading of epidermal cells / negative regulation of type I interferon-mediated signaling pathway / lung alveolus development ...macrophage elastase / negative regulation of endothelial cell-matrix adhesion / bronchiole development / positive regulation of epithelial cell proliferation involved in wound healing / elastin catabolic process / regulation of defense response to virus by host / positive regulation of type I interferon-mediated signaling pathway / wound healing, spreading of epidermal cells / negative regulation of type I interferon-mediated signaling pathway / lung alveolus development / response to amyloid-beta / Collagen degradation / collagen catabolic process / positive regulation of interferon-alpha production / extracellular matrix disassembly / core promoter sequence-specific DNA binding / collagen binding / Degradation of the extracellular matrix / extracellular matrix organization / metalloendopeptidase activity / cellular response to virus / protein import into nucleus / extracellular matrix / endopeptidase activity / sequence-specific DNA binding / serine-type endopeptidase activity / calcium ion binding / negative regulation of transcription by RNA polymerase II / positive regulation of transcription by RNA polymerase II / proteolysis / : / extracellular region / zinc ion binding / nucleus / cytoplasm Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.8 Å MOLECULAR REPLACEMENT / Resolution: 1.8 Å | |||||||||
 Authors Authors | Tepshi, L. / Bordenave, T. / Rouanet-Mehouas, C. / Devel, L. / Dive, V. / Stura, E.A. | |||||||||
 Citation Citation |  Journal: Bioconjug.Chem. / Year: 2016 Journal: Bioconjug.Chem. / Year: 2016Title: Synthesis and in Vitro and in Vivo Evaluation of MMP-12 Selective Optical Probes. Authors: Bordenave, T. / Helle, M. / Beau, F. / Georgiadis, D. / Tepshi, L. / Bernes, M. / Ye, Y. / Levenez, L. / Poquet, E. / Nozach, H. / Razavian, M. / Toczek, J. / Stura, E.A. / Dive, V. / Sadeghi, M.M. / Devel, L. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  5l7f.cif.gz 5l7f.cif.gz | 102.3 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb5l7f.ent.gz pdb5l7f.ent.gz | 75.2 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  5l7f.json.gz 5l7f.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/l7/5l7f https://data.pdbj.org/pub/pdb/validation_reports/l7/5l7f ftp://data.pdbj.org/pub/pdb/validation_reports/l7/5l7f ftp://data.pdbj.org/pub/pdb/validation_reports/l7/5l7f | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  5l79C  5czmS S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 | 
| ||||||||
| Unit cell |
|
- Components
Components
-Protein , 1 types, 2 molecules AB
| #1: Protein | Mass: 17557.568 Da / Num. of mol.: 2 / Mutation: F171D K241A Source method: isolated from a genetically manipulated source Details: Fragment: catalytic domain (UNP residues 106-263) mutations: F171D K241A Source: (gene. exp.)  Homo sapiens (human) / Gene: MMP12, HME / Production host: Homo sapiens (human) / Gene: MMP12, HME / Production host:  |
|---|
-Non-polymers , 9 types, 384 molecules 




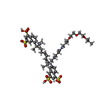







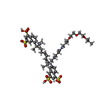



| #2: Chemical | ChemComp-ZN / #3: Chemical | ChemComp-CA / #4: Chemical | ChemComp-BR / | #5: Chemical | #6: Chemical |   Type: peptide-like, Peptide-like / Class: Inhibitor / Mass: 818.004 Da / Num. of mol.: 2 / Source method: obtained synthetically / Formula: C35H35BrClN4O10P Type: peptide-like, Peptide-like / Class: Inhibitor / Mass: 818.004 Da / Num. of mol.: 2 / Source method: obtained synthetically / Formula: C35H35BrClN4O10PReferences: N-[(2S)-3-[(S)-(4-BROMOPHENYL)(HYDROXY)PHOSPHORYL]-2-{[3-(3'-CHLOROBIPHENYL-4-YL)-1,2-OXAZOL-5-YL]METHYL}PROPANOYL]-L-ALPHA-GLUTAMYL-L-ALPHA-GLUTAMINE #7: Chemical | #8: Chemical | ChemComp-GOL / | #9: Chemical | ChemComp-DMS / | #10: Water | ChemComp-HOH / | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.48 Å3/Da / Density % sol: 50.34 % / Description: flat plate |
|---|---|
| Crystal grow | Temperature: 293 K / Method: vapor diffusion, sitting drop / pH: 9.5 Details: protein: MMP12-F171D-K241A 661 micro-M + 10 milli-M AHA, 10% DMSO, 0.5 milli-M fluorescent inhibitor (R47-CY5) precipitant: 20% PEG4K, 2% gamma valerolactone, 0.2 milli-M TRIS pH 9.5 40% ...Details: protein: MMP12-F171D-K241A 661 micro-M + 10 milli-M AHA, 10% DMSO, 0.5 milli-M fluorescent inhibitor (R47-CY5) precipitant: 20% PEG4K, 2% gamma valerolactone, 0.2 milli-M TRIS pH 9.5 40% CM26 ((12.5 % diethylene glycol + 12.5 % ethylene glycol + 12.5 % glycerol + 25 % 2,3-butanediol + 12.5 % DMSO), 25% MPEG 6K PH range: 9.5 / Temp details: cooled incubator |
-Data collection
| Diffraction | Mean temperature: 100 K / Ambient temp details: cryostream |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  SOLEIL SOLEIL  / Beamline: PROXIMA 1 / Wavelength: 0.97857 Å / Beamline: PROXIMA 1 / Wavelength: 0.97857 Å |
| Detector | Type: DECTRIS PILATUS 6M / Detector: PIXEL / Date: Dec 10, 2015 / Details: mirrors |
| Radiation | Monochromator: crystal / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.97857 Å / Relative weight: 1 |
| Reflection | Resolution: 1.79→47.34 Å / Num. obs: 63601 / % possible obs: 96.9 % / Observed criterion σ(F): 0 / Observed criterion σ(I): -3 / Redundancy: 2.386860603 % / CC1/2: 0.988 / Rmerge(I) obs: 0.145 / Rsym value: 0.113 / Net I/σ(I): 6.87 |
| Reflection shell | Resolution: 1.79→1.84 Å / Redundancy: 2.36172043 % / Rmerge(I) obs: 0.947 / Mean I/σ(I) obs: 1.43 / % possible all: 98.4 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 5CZM Resolution: 1.8→47.34 Å / Cor.coef. Fo:Fc: 0.959 / Cor.coef. Fo:Fc free: 0.951 / SU B: 3.324 / SU ML: 0.097 / Cross valid method: THROUGHOUT / ESU R: 0.139 / ESU R Free: 0.117 / Details: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Ion probe radii: 0.8 Å / Shrinkage radii: 0.8 Å / VDW probe radii: 1.2 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 21.924 Å2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: 1 / Resolution: 1.8→47.34 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
|
 Movie
Movie Controller
Controller











 PDBj
PDBj












