[English] 日本語
 Yorodumi
Yorodumi- PDB-5l5n: Plexin A4 full extracellular region, domains 1 to 7 modeled, data... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 5l5n | ||||||
|---|---|---|---|---|---|---|---|
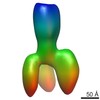
| Title | Plexin A4 full extracellular region, domains 1 to 7 modeled, data to 8.5 angstrom, spacegroup P4(3)22 | ||||||
 Components Components | Plexin-A4 | ||||||
 Keywords Keywords | SIGNALING PROTEIN / receptor / signaling / axon guidance | ||||||
| Function / homology |  Function and homology information Function and homology informationglossopharyngeal nerve morphogenesis / chemorepulsion of branchiomotor axon / regulation of negative chemotaxis / vagus nerve morphogenesis / cranial nerve morphogenesis / trigeminal nerve morphogenesis / anterior commissure morphogenesis / regulation of axon extension involved in axon guidance / postganglionic parasympathetic fiber development / facial nerve morphogenesis ...glossopharyngeal nerve morphogenesis / chemorepulsion of branchiomotor axon / regulation of negative chemotaxis / vagus nerve morphogenesis / cranial nerve morphogenesis / trigeminal nerve morphogenesis / anterior commissure morphogenesis / regulation of axon extension involved in axon guidance / postganglionic parasympathetic fiber development / facial nerve morphogenesis / CRMPs in Sema3A signaling / Sema3A PAK dependent Axon repulsion / sympathetic neuron axon guidance / trigeminal nerve structural organization / facial nerve structural organization / branchiomotor neuron axon guidance / SEMA3A-Plexin repulsion signaling by inhibiting Integrin adhesion / maintenance of synapse structure / cerebellar climbing fiber to Purkinje cell synapse / semaphorin receptor complex / sympathetic nervous system development / semaphorin receptor activity / embryonic heart tube development / motor neuron axon guidance / semaphorin-plexin signaling pathway / neuron projection morphogenesis / synapse assembly / regulation of cell migration / axon guidance / nervous system development / plasma membrane Similarity search - Function | ||||||
| Biological species |  | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 8.502 Å MOLECULAR REPLACEMENT / Resolution: 8.502 Å | ||||||
 Authors Authors | Janssen, B.J.C. / Kong, Y. / Malinauskas, T. / Vangoor, V.R. / Coles, C.H. / Kaufmann, R. / Ni, T. / Gilbert, R.J.C. / Padilla-Parra, S. / Pasterkamp, R.J. / Jones, E.Y. | ||||||
 Citation Citation |  Journal: Neuron / Year: 2016 Journal: Neuron / Year: 2016Title: Structural Basis for Plexin Activation and Regulation. Authors: Kong, Y. / Janssen, B.J. / Malinauskas, T. / Vangoor, V.R. / Coles, C.H. / Kaufmann, R. / Ni, T. / Gilbert, R.J. / Padilla-Parra, S. / Pasterkamp, R.J. / Jones, E.Y. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  5l5n.cif.gz 5l5n.cif.gz | 179.6 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb5l5n.ent.gz pdb5l5n.ent.gz | 130 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  5l5n.json.gz 5l5n.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/l5/5l5n https://data.pdbj.org/pub/pdb/validation_reports/l5/5l5n ftp://data.pdbj.org/pub/pdb/validation_reports/l5/5l5n ftp://data.pdbj.org/pub/pdb/validation_reports/l5/5l5n | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  5l56SC  5l59C  5l5cC  5l5gC  5l5kC  5l5lC  5l5mC  5l74C  5l7nC S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 134813.844 Da / Num. of mol.: 1 / Fragment: UNP residues 36-1229 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Homo sapiens (human) / References: UniProt: Q80UG2 Homo sapiens (human) / References: UniProt: Q80UG2 |
|---|---|
| Has protein modification | Y |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density % sol: 83 % |
|---|---|
| Crystal grow | Temperature: 293.5 K / Method: vapor diffusion, sitting drop / pH: 8.5 / Details: 1.5 lithium sulfate, 100 mM TRIS, pH 8.5 |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  Diamond Diamond  / Beamline: I04 / Wavelength: 0.9763 Å / Beamline: I04 / Wavelength: 0.9763 Å |
| Detector | Type: ADSC QUANTUM 315 / Detector: CCD / Date: Oct 22, 2010 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.9763 Å / Relative weight: 1 |
| Reflection | Resolution: 8.5→71 Å / Num. obs: 4374 / % possible obs: 99.7 % / Redundancy: 5.8 % / Rmerge(I) obs: 0.33 / Net I/σ(I): 4.2 |
| Reflection shell | Highest resolution: 8.5 Å / Rmerge(I) obs: 1.17 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 5L56 Resolution: 8.502→70.367 Å / SU ML: 1.89 / Cross valid method: FREE R-VALUE / σ(F): 1.12 / Phase error: 45.03 Details: THE AUTHORS STATE THAT DUE TO THE LOW RESOLUTION OF THE DIFFRACTION DATA THE STRUCTURE WAS ONLY SUBJECTED TO RIGID BODY REFINEMENT WITH EACH DOMAIN AS RIGID BODY. THE STRUCTURE IS BASED ON A ...Details: THE AUTHORS STATE THAT DUE TO THE LOW RESOLUTION OF THE DIFFRACTION DATA THE STRUCTURE WAS ONLY SUBJECTED TO RIGID BODY REFINEMENT WITH EACH DOMAIN AS RIGID BODY. THE STRUCTURE IS BASED ON A HOMOLOGY MODEL GENERATED WITH PDB ENTRY 5L56 AS TEMPLATE (53% SEQUENCE IDENTITY). THE AUTHORS HAVE NOT FURTHER REFINED THE RESULTING COORDINATES NOR CORRECTED RESULTING CLASHES BETWEEN ATOMS AND DEVIATING PEPTIDE LINKAGES BETWEEN DOMAINS.
| ||||||||||||||||||||||||||||
| Solvent computation | Shrinkage radii: 0.9 Å / VDW probe radii: 1.11 Å | ||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 8.502→70.367 Å
| ||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||
| LS refinement shell |
|
 Movie
Movie Controller
Controller






 PDBj
PDBj


