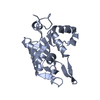
Entry Database : PDB / ID : 5h9wTitle Crystal structure of Regnase PIN domain, form II Ribonuclease ZC3H12A Keywords / Function / homology Function Domain/homology Component
/ / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / Biological species Mus musculus (house mouse)Method / / / Resolution : 2.6 Å Authors Yokogawa, M. / Tsushima, T. / Adachi, W. / Noda, N.N. / Inagaki, F. Funding support Organization Grant number Country JSPS
Journal : Sci Rep / Year : 2016Title : Structural basis for the regulation of enzymatic activity of Regnase-1 by domain-domain interactionsAuthors : Yokogawa, M. / Tsushima, T. / Noda, N.N. / Kumeta, H. / Enokizono, Y. / Yamashita, K. / Standley, D.M. / Takeuchi, O. / Akira, S. / Inagaki, F. History Deposition Dec 29, 2015 Deposition site / Processing site Revision 1.0 Mar 16, 2016 Provider / Type Revision 1.1 Feb 19, 2020 Group / Derived calculations / Category / pdbx_struct_oper_listItem / _pdbx_struct_oper_list.symmetry_operationRevision 1.2 Nov 8, 2023 Group / Database references / Refinement descriptionCategory chem_comp_atom / chem_comp_bond ... chem_comp_atom / chem_comp_bond / database_2 / pdbx_initial_refinement_model / struct_ncs_dom_lim Item _database_2.pdbx_DOI / _database_2.pdbx_database_accession ... _database_2.pdbx_DOI / _database_2.pdbx_database_accession / _struct_ncs_dom_lim.beg_auth_comp_id / _struct_ncs_dom_lim.beg_label_asym_id / _struct_ncs_dom_lim.beg_label_comp_id / _struct_ncs_dom_lim.beg_label_seq_id / _struct_ncs_dom_lim.end_auth_comp_id / _struct_ncs_dom_lim.end_label_asym_id / _struct_ncs_dom_lim.end_label_comp_id / _struct_ncs_dom_lim.end_label_seq_id
Show all Show less
 Open data
Open data Basic information
Basic information Components
Components Keywords
Keywords Function and homology information
Function and homology information
 X-RAY DIFFRACTION /
X-RAY DIFFRACTION /  SYNCHROTRON /
SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2.6 Å
MOLECULAR REPLACEMENT / Resolution: 2.6 Å  Authors
Authors Japan, 1items
Japan, 1items  Citation
Citation Journal: Sci Rep / Year: 2016
Journal: Sci Rep / Year: 2016 Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 5h9w.cif.gz
5h9w.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb5h9w.ent.gz
pdb5h9w.ent.gz PDB format
PDB format 5h9w.json.gz
5h9w.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads https://data.pdbj.org/pub/pdb/validation_reports/h9/5h9w
https://data.pdbj.org/pub/pdb/validation_reports/h9/5h9w ftp://data.pdbj.org/pub/pdb/validation_reports/h9/5h9w
ftp://data.pdbj.org/pub/pdb/validation_reports/h9/5h9w



 Links
Links Assembly
Assembly


 Components
Components

 X-RAY DIFFRACTION
X-RAY DIFFRACTION Sample preparation
Sample preparation SYNCHROTRON / Site:
SYNCHROTRON / Site:  SPring-8
SPring-8  / Beamline: BL41XU / Wavelength: 1 Å
/ Beamline: BL41XU / Wavelength: 1 Å Processing
Processing MOLECULAR REPLACEMENT
MOLECULAR REPLACEMENT Movie
Movie Controller
Controller












 PDBj
PDBj

