+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 5h48 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Crystal structure of Cbln1 | |||||||||
 Components Components | Cerebellin-1 | |||||||||
 Keywords Keywords | PROTEIN BINDING / C1q domain / C1q/TNF superfamily / receptor specificity / synaptogenesis | |||||||||
| Function / homology |  Function and homology information Function and homology informationnegative regulation of inhibitory synapse assembly / trans-synaptic signaling, modulating synaptic transmission / trans-synaptic protein complex / cerebellar granule cell differentiation / positive regulation of long-term synaptic depression / maintenance of synapse structure / regulation of postsynaptic density assembly / negative regulation of excitatory postsynaptic potential / positive regulation of synapse assembly / heterophilic cell-cell adhesion ...negative regulation of inhibitory synapse assembly / trans-synaptic signaling, modulating synaptic transmission / trans-synaptic protein complex / cerebellar granule cell differentiation / positive regulation of long-term synaptic depression / maintenance of synapse structure / regulation of postsynaptic density assembly / negative regulation of excitatory postsynaptic potential / positive regulation of synapse assembly / heterophilic cell-cell adhesion / parallel fiber to Purkinje cell synapse / protein secretion / regulation of presynapse assembly / synaptic cleft / synapse assembly / establishment of localization in cell / synapse organization / postsynaptic membrane / synapse / glutamatergic synapse / extracellular region / identical protein binding Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2.2 Å MOLECULAR REPLACEMENT / Resolution: 2.2 Å | |||||||||
 Authors Authors | Zhong, C. / Shen, J. / Zhang, H. / Ding, J. | |||||||||
| Funding support |  China, 2items China, 2items
| |||||||||
 Citation Citation |  Journal: Cell Rep / Year: 2017 Journal: Cell Rep / Year: 2017Title: Cbln1 and Cbln4 Are Structurally Similar but Differ in GluD2 Binding Interactions. Authors: Chen Zhong / Jinlong Shen / Huibing Zhang / Guangyi Li / Senlin Shen / Fang Wang / Kuan Hu / Longxing Cao / Yongning He / Jianping Ding /  Abstract: Unlike cerebellin 1 (Cbln1), which bridges neurexin (Nrxn) receptors and δ-type glutamate receptors in a trans-synaptic triad, Cbln4 was reported to have no or weak binding for the receptors ...Unlike cerebellin 1 (Cbln1), which bridges neurexin (Nrxn) receptors and δ-type glutamate receptors in a trans-synaptic triad, Cbln4 was reported to have no or weak binding for the receptors despite sharing ∼70% sequence identity with Cbln1. Here, we report crystal structures of the homotrimers of the C1q domain of Cbln1 and Cbln4 at 2.2 and 2.3 Å resolution, respectively. Comparison of the structures suggests that the difference between Cbln1 and Cbln4 in GluD2 binding might be because of their sequence and structural divergence in loop CD. Surprisingly, we show that Cbln4 binds to Nrxn1β and forms a stable complex with the laminin, nectin, sex-hormone binding globulin (LNS) domain of Nrxn1β. Furthermore, the negative-stain electron microscopy reconstruction of hexameric full-length Cbln1 at 13 Å resolution and that of the Cbln4/Nrxn1β complex at 19 Å resolution suggest that Nrxn1β binds to the N-terminal region of Cbln4, probably through strand β10 of the S4 insert. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  5h48.cif.gz 5h48.cif.gz | 46 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb5h48.ent.gz pdb5h48.ent.gz | 29.6 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  5h48.json.gz 5h48.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/h4/5h48 https://data.pdbj.org/pub/pdb/validation_reports/h4/5h48 ftp://data.pdbj.org/pub/pdb/validation_reports/h4/5h48 ftp://data.pdbj.org/pub/pdb/validation_reports/h4/5h48 | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  6665C 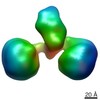 6666C  5h49C  5h4bC  5h4cC  4nn0S C: citing same article ( S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 20418.984 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Homo sapiens (human) / References: UniProt: P63182 Homo sapiens (human) / References: UniProt: P63182 |
|---|---|
| #2: Polysaccharide | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose Source method: isolated from a genetically manipulated source |
| #3: Water | ChemComp-HOH / |
| Has protein modification | Y |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.46 Å3/Da / Density % sol: 50 % |
|---|---|
| Crystal grow | Temperature: 298 K / Method: vapor diffusion, hanging drop Details: 0.2 M Sodium tartrate dibasic dihydrate, 20% PEG3350, pH 7.3 |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  SSRF SSRF  / Beamline: BL17U1 / Wavelength: 0.9199 Å / Beamline: BL17U1 / Wavelength: 0.9199 Å |
| Detector | Type: RAYONIX MX-325 / Detector: CCD / Date: Oct 31, 2014 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.9199 Å / Relative weight: 1 |
| Reflection | Resolution: 2.2→71.86 Å / Num. obs: 9670 / % possible obs: 94.7 % / Redundancy: 3.9 % / Net I/σ(I): 13.1 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 4NN0 Resolution: 2.2→71.86 Å / Cor.coef. Fo:Fc: 0.963 / Cor.coef. Fo:Fc free: 0.937 / SU B: 4.473 / SU ML: 0.112 / Cross valid method: THROUGHOUT / ESU R: 0.192 / ESU R Free: 0.166 / Stereochemistry target values: MAXIMUM LIKELIHOOD / Details: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Ion probe radii: 0.8 Å / Shrinkage radii: 0.8 Å / VDW probe radii: 1.2 Å / Solvent model: MASK | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 35.338 Å2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: 1 / Resolution: 2.2→71.86 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
|
 Movie
Movie Controller
Controller












 PDBj
PDBj