[English] 日本語
 Yorodumi
Yorodumi- PDB-4s2q: Crystal Structure of HMG domain of the chondrogenesis master regu... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 4s2q | ||||||
|---|---|---|---|---|---|---|---|
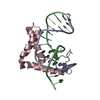
| Title | Crystal Structure of HMG domain of the chondrogenesis master regulator, Sox9 in complex with ChIP-Seq identified DNA element | ||||||
 Components Components |
| ||||||
 Keywords Keywords | DNA BINDING PROTEIN/DNA / DNA Bending / Minor groove binding / Transcription Regulation / DNA BINDING PROTEIN-DNA complex | ||||||
| Function / homology |  Function and homology information Function and homology informationheart valve formation / male germ-line sex determination / epithelial cell proliferation involved in prostatic bud elongation / regulation of cell proliferation involved in tissue homeostasis / regulation of branching involved in lung morphogenesis / morphogenesis of a branching epithelium / cell proliferation involved in heart morphogenesis / renal vesicle induction / metanephric tubule development / ureter urothelium development ...heart valve formation / male germ-line sex determination / epithelial cell proliferation involved in prostatic bud elongation / regulation of cell proliferation involved in tissue homeostasis / regulation of branching involved in lung morphogenesis / morphogenesis of a branching epithelium / cell proliferation involved in heart morphogenesis / renal vesicle induction / metanephric tubule development / ureter urothelium development / positive regulation of kidney development / negative regulation of beta-catenin-TCF complex assembly / positive regulation of mesenchymal stem cell differentiation / regulation of epithelial cell proliferation involved in lung morphogenesis / neural crest cell fate specification / ureter smooth muscle cell differentiation / metanephric nephron tubule formation / negative regulation of immune system process / : / glial cell fate specification / intrahepatic bile duct development / astrocyte fate commitment / bronchus cartilage development / lung smooth muscle development / ureter development / ureter morphogenesis / chondrocyte differentiation involved in endochondral bone morphogenesis / Transcriptional regulation by RUNX2 / negative regulation of fatty acid oxidation / anterior head development / chondrocyte hypertrophy / Harderian gland development / cellular response to heparin / Sertoli cell differentiation / otic vesicle development / trachea cartilage development / retinal rod cell differentiation / growth plate cartilage chondrocyte growth / chondrocyte development / positive regulation of cell proliferation involved in heart morphogenesis / negative regulation of photoreceptor cell differentiation / intestinal epithelial cell differentiation / positive regulation of epithelial cell differentiation / regulation of cell cycle process / glandular epithelial cell differentiation / lacrimal gland development / prostate gland morphogenesis / otic vesicle formation / positive regulation of male gonad development / Deactivation of the beta-catenin transactivating complex / extracellular matrix assembly / endochondral bone morphogenesis / negative regulation of mesenchymal cell apoptotic process / positive regulation of cartilage development / bHLH transcription factor binding / neuron fate specification / positive regulation of chondrocyte differentiation / prostate gland development / notochord development / limb bud formation / positive regulation of extracellular matrix assembly / heart valve morphogenesis / mesenchymal cell apoptotic process / neural crest cell development / lung epithelial cell differentiation / Sertoli cell development / cochlea morphogenesis / negative regulation of bone mineralization / endocrine pancreas development / mesenchymal cell proliferation / intestinal epithelial structure maintenance / response to fatty acid / positive regulation of chondrocyte proliferation / positive regulation of branching involved in ureteric bud morphogenesis / tissue homeostasis / cellular response to BMP stimulus / negative regulation of biomineral tissue development / male sex determination / heart valve development / negative regulation of myoblast differentiation / mammary gland development / cartilage development / negative regulation of chondrocyte differentiation / negative regulation of ossification / aortic valve morphogenesis / positive regulation of mesenchymal cell proliferation / endocardial cushion morphogenesis / branching involved in ureteric bud morphogenesis / cartilage condensation / negative regulation of epithelial cell differentiation / positive regulation of stem cell proliferation / epithelial tube branching involved in lung morphogenesis / bone mineralization / protein kinase A catalytic subunit binding / type I pneumocyte differentiation / regulation of cell differentiation / oligodendrocyte differentiation / positive regulation of epithelial cell migration / epithelial to mesenchymal transition / cellular response to interleukin-1 Similarity search - Function | ||||||
| Biological species |  synthetic construct (others) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2.7 Å MOLECULAR REPLACEMENT / Resolution: 2.7 Å | ||||||
 Authors Authors | Vivekanandan, S. / Moovarkumudalvan, B. / Lescar, J. / Kolatkar, P.R. | ||||||
 Citation Citation |  Journal: To be Published Journal: To be PublishedTitle: Crystal Structure of HMG domain of the chondrogenesis master regulator, Sox9 in complex with ChIP-Seq identified DNA element Authors: Vivekanandan, S. / Moovarkumudalvan, B. / Lescar, J. / Kolatkar, P.R. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  4s2q.cif.gz 4s2q.cif.gz | 83.4 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb4s2q.ent.gz pdb4s2q.ent.gz | 60.7 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  4s2q.json.gz 4s2q.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Summary document |  4s2q_validation.pdf.gz 4s2q_validation.pdf.gz | 431.9 KB | Display |  wwPDB validaton report wwPDB validaton report |
|---|---|---|---|---|
| Full document |  4s2q_full_validation.pdf.gz 4s2q_full_validation.pdf.gz | 433.6 KB | Display | |
| Data in XML |  4s2q_validation.xml.gz 4s2q_validation.xml.gz | 6.3 KB | Display | |
| Data in CIF |  4s2q_validation.cif.gz 4s2q_validation.cif.gz | 7.6 KB | Display | |
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/s2/4s2q https://data.pdbj.org/pub/pdb/validation_reports/s2/4s2q ftp://data.pdbj.org/pub/pdb/validation_reports/s2/4s2q ftp://data.pdbj.org/pub/pdb/validation_reports/s2/4s2q | HTTPS FTP |
-Related structure data
| Related structure data |  3f27S S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: DNA chain | Mass: 4871.149 Da / Num. of mol.: 1 / Source method: obtained synthetically / Source: (synth.) synthetic construct (others) |
|---|---|
| #2: DNA chain | Mass: 4965.257 Da / Num. of mol.: 1 / Source method: obtained synthetically / Source: (synth.) synthetic construct (others) |
| #3: Protein | Mass: 9264.711 Da / Num. of mol.: 1 / Fragment: UNP residues 103-178 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   |
| #4: Water | ChemComp-HOH / |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.97 Å3/Da / Density % sol: 58.63 % |
|---|---|
| Crystal grow | Temperature: 291 K / Method: vapor diffusion, hanging drop / pH: 8.5 Details: 20% PEG 3350, 200mM Sodium/potassium phosphate, 100mM Bis Tris propane, pH 8.5 , VAPOR DIFFUSION, HANGING DROP, temperature 291K |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  NSLS NSLS  / Beamline: X29A / Wavelength: 1.075 Å / Beamline: X29A / Wavelength: 1.075 Å |
| Detector | Type: ADSC QUANTUM 315 / Detector: CCD / Date: Aug 10, 2012 |
| Radiation | Monochromator: ROSENBAUM-ROCK DOUBLE CRYSTAL SAGITTAL FOCUSING MONOCHROMATOR Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.075 Å / Relative weight: 1 |
| Reflection | Resolution: 2.7→50 Å / Num. all: 6721 / Num. obs: 6675 / % possible obs: 99.9 % / Observed criterion σ(F): 0 / Observed criterion σ(I): 2 / Redundancy: 30.3 % / Rsym value: 0.133 / Net I/σ(I): 19 |
| Reflection shell | Resolution: 2.7→3.4011 Å / Redundancy: 30.3 % / Mean I/σ(I) obs: 19 / Num. unique all: 6721 / Rsym value: 0.133 / % possible all: 99.9 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB entry 3F27 Resolution: 2.7→24.873 Å / SU ML: 0.23 / σ(F): 1.34 / Phase error: 32.56 / Stereochemistry target values: ML
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Shrinkage radii: 0.9 Å / VDW probe radii: 1.11 Å / Solvent model: FLAT BULK SOLVENT MODEL | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.7→24.873 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS params. | Method: refined / Refine-ID: X-RAY DIFFRACTION
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS group |
|
 Movie
Movie Controller
Controller









 PDBj
PDBj








































