Entry Database : PDB / ID : 4p9vTitle Grb2 SH2 complexed with a pTyr-Ac6cN-Asn tripeptide Growth factor receptor-bound protein 2 PHQ-PTR-02K-ASN-NH2 Keywords / / / Function / homology Function Domain/homology Component
/ / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / Biological species Homo sapiens (human)synthetic construct (others) Method / / Resolution : 1.64 Å Authors Clements, J.H. / Martin, S.F. Funding support Organization Grant number Country National Institutes of Health/National Institute of General Medical Sciences (NIH/NIGMS) GM 84965 National Science Foundation (NSF, United States) CHE 0750329 Robert A. Welch Foundation F-652
Journal : Bioorg.Med.Chem.Lett. / Year : 2014Title : Protein-ligand interactions: Probing the energetics of a putative cation-pi interaction.Authors : Myslinski, J.M. / Clements, J.H. / Martin, S.F. History Deposition Apr 6, 2014 Deposition site / Processing site Revision 1.0 Jun 18, 2014 Provider / Type Revision 1.1 Jul 16, 2014 Group Revision 1.2 Dec 24, 2014 Group Revision 1.3 Aug 9, 2017 Group Database references / Derived calculations ... Database references / Derived calculations / Other / Source and taxonomy Category citation / entity_src_gen ... citation / entity_src_gen / pdbx_database_status / pdbx_entity_src_syn / pdbx_struct_oper_list Item _citation.journal_id_CSD / _entity_src_gen.pdbx_alt_source_flag ... _citation.journal_id_CSD / _entity_src_gen.pdbx_alt_source_flag / _pdbx_database_status.pdb_format_compatible / _pdbx_entity_src_syn.pdbx_alt_source_flag / _pdbx_struct_oper_list.symmetry_operation Revision 1.4 Sep 27, 2017 Group / Category / Item Revision 1.5 Nov 27, 2019 Group / Category / Item Revision 1.6 Sep 27, 2023 Group / Database references / Refinement descriptionCategory chem_comp_atom / chem_comp_bond ... chem_comp_atom / chem_comp_bond / database_2 / pdbx_initial_refinement_model / refine_hist Item _database_2.pdbx_DOI / _database_2.pdbx_database_accession ... _database_2.pdbx_DOI / _database_2.pdbx_database_accession / _refine_hist.number_atoms_solvent / _refine_hist.number_atoms_total / _refine_hist.pdbx_number_atoms_ligand / _refine_hist.pdbx_number_atoms_nucleic_acid / _refine_hist.pdbx_number_atoms_protein Revision 1.7 Nov 15, 2023 Group / Category / chem_comp_bond / Item / _chem_comp_bond.atom_id_2Revision 1.8 Nov 13, 2024 Group / Category / pdbx_modification_feature
Show all Show less
 Open data
Open data Basic information
Basic information Components
Components Keywords
Keywords Function and homology information
Function and homology information Homo sapiens (human)
Homo sapiens (human) X-RAY DIFFRACTION /
X-RAY DIFFRACTION /  MOLECULAR REPLACEMENT / Resolution: 1.64 Å
MOLECULAR REPLACEMENT / Resolution: 1.64 Å  Authors
Authors United States, 3items
United States, 3items  Citation
Citation Journal: Bioorg.Med.Chem.Lett. / Year: 2014
Journal: Bioorg.Med.Chem.Lett. / Year: 2014 Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 4p9v.cif.gz
4p9v.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb4p9v.ent.gz
pdb4p9v.ent.gz PDB format
PDB format 4p9v.json.gz
4p9v.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads https://data.pdbj.org/pub/pdb/validation_reports/p9/4p9v
https://data.pdbj.org/pub/pdb/validation_reports/p9/4p9v ftp://data.pdbj.org/pub/pdb/validation_reports/p9/4p9v
ftp://data.pdbj.org/pub/pdb/validation_reports/p9/4p9v
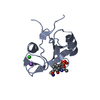
 Links
Links Assembly
Assembly
 Components
Components Homo sapiens (human) / Gene: GRB2, ASH / Plasmid: pQE-60 / Production host:
Homo sapiens (human) / Gene: GRB2, ASH / Plasmid: pQE-60 / Production host: 
 X-RAY DIFFRACTION / Number of used crystals: 1
X-RAY DIFFRACTION / Number of used crystals: 1  Sample preparation
Sample preparation ROTATING ANODE / Type: RIGAKU RU200 / Wavelength: 1.5418 Å
ROTATING ANODE / Type: RIGAKU RU200 / Wavelength: 1.5418 Å Processing
Processing MOLECULAR REPLACEMENT
MOLECULAR REPLACEMENT Movie
Movie Controller
Controller








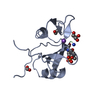

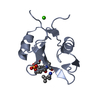


 PDBj
PDBj

























