[English] 日本語
 Yorodumi
Yorodumi- PDB-4lea: The Crystal Structure of Pyocin L1 bound to D-mannose at 2.55 Ang... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 4lea | ||||||
|---|---|---|---|---|---|---|---|
| Title | The Crystal Structure of Pyocin L1 bound to D-mannose at 2.55 Angstroms | ||||||
 Components Components | Pyocin L1 | ||||||
 Keywords Keywords | SUGAR BINDING PROTEIN / Mannose Monocot Binding Protein / MMBL / Galanthus nivalis Agglutinin / Beta Prism / Protein Antibiotic / Extra cellular | ||||||
| Function / homology |  Function and homology information Function and homology informationAgglutinin, subunit A - #30 / Agglutinin, subunit A / Bulb-type lectin domain / Bulb-type lectin domain / Bulb-type lectin domain superfamily / Bulb-type lectin domain profile. / Bulb-type mannose-specific lectin / Orthogonal Prism / Mainly Beta Similarity search - Domain/homology | ||||||
| Biological species |  | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2.55 Å MOLECULAR REPLACEMENT / Resolution: 2.55 Å | ||||||
 Authors Authors | Grinter, R. / Roszak, A.W. / Mccaughey, L. / Cogdell, C.J. / Walker, D. | ||||||
 Citation Citation |  Journal: Plos Pathog. / Year: 2014 Journal: Plos Pathog. / Year: 2014Title: Lectin-Like Bacteriocins from Pseudomonas spp. Utilise D-Rhamnose Containing Lipopolysaccharide as a Cellular Receptor. Authors: McCaughey, L.C. / Grinter, R. / Josts, I. / Roszak, A.W. / Walen, K.I. / Cogdell, R.J. / Milner, J. / Evans, T. / Kelly, S. / Tucker, N.P. / Byron, O. / Smith, B. / Walker, D. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  4lea.cif.gz 4lea.cif.gz | 113.3 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb4lea.ent.gz pdb4lea.ent.gz | 88.9 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  4lea.json.gz 4lea.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/le/4lea https://data.pdbj.org/pub/pdb/validation_reports/le/4lea ftp://data.pdbj.org/pub/pdb/validation_reports/le/4lea ftp://data.pdbj.org/pub/pdb/validation_reports/le/4lea | HTTPS FTP |
|---|
-Related structure data
| Related structure data | 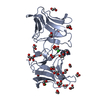 4le7SC  4ledC S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 | 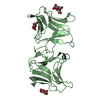
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 30008.020 Da / Num. of mol.: 2 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   #2: Sugar | ChemComp-BMA / #3: Water | ChemComp-HOH / | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.75 Å3/Da / Density % sol: 55.27 % |
|---|---|
| Crystal grow | Temperature: 293 K / Method: vapor diffusion, sitting drop / pH: 7.5 Details: 20% PEG 550 MME, 20% PEG 20K, 0.03 M CaCl2, 0.03 M MgCl2 0.1 M MOPS/HEPES, pH 7.5, VAPOR DIFFUSION, SITTING DROP, temperature 293K |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  Diamond Diamond  / Beamline: I24 / Wavelength: 0.9762 Å / Beamline: I24 / Wavelength: 0.9762 Å |
| Detector | Type: PSI PILATUS 6M / Detector: PIXEL / Date: Dec 14, 2012 |
| Radiation | Monochromator: Silicon / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.9762 Å / Relative weight: 1 |
| Reflection | Resolution: 2.55→55.53 Å / Num. obs: 22096 / % possible obs: 99.9 % / Observed criterion σ(F): 2 / Observed criterion σ(I): 2 / Redundancy: 5.5 % / Biso Wilson estimate: 61.517 Å2 / Rmerge(I) obs: 0.059 / Net I/σ(I): 13.3 |
| Reflection shell | Resolution: 2.55→2.67 Å / % possible all: 99.5 |
- Processing
Processing
| Software |
| ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB ENTRY 4LE7 Resolution: 2.55→42.28 Å / σ(F): 2 / Stereochemistry target values: MAXIMUM LIKELIHOOD
| ||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.55→42.28 Å
|
 Movie
Movie Controller
Controller












 PDBj
PDBj



