+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 430d | ||||||
|---|---|---|---|---|---|---|---|
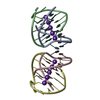
| Title | STRUCTURE OF SARCIN/RICIN LOOP FROM RAT 28S RRNA | ||||||
 Components Components | SARCIN/RICIN LOOP FROM RAT 28S R-RNA | ||||||
 Keywords Keywords | RNA / U-RNA / DOUBLE HELIX / HAIRPIN / BLUNT STEM / TETRALOOP MISMATCHED / BASE TRIPLE / RIBONUCLEIC ACID | ||||||
| Function / homology | RNA / RNA (> 10) Function and homology information Function and homology information | ||||||
| Biological species |  | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / MAD PHASING METHOD / Resolution: 2.1 Å SYNCHROTRON / MAD PHASING METHOD / Resolution: 2.1 Å | ||||||
 Authors Authors | Correll, C.C. / Munishkin, A. / Chan, Y.L. / Ren, Z. / Wool, I.G. / Steitz, T.A. | ||||||
 Citation Citation |  Journal: Proc.Natl.Acad.Sci.USA / Year: 1998 Journal: Proc.Natl.Acad.Sci.USA / Year: 1998Title: Crystal structure of the ribosomal RNA domain essential for binding elongation factors. Authors: Correll, C.C. / Munishkin, A. / Chan, Y.L. / Ren, Z. / Wool, I.G. / Steitz, T.A. #1:  Journal: Proc.Natl.Acad.Sci.USA / Year: 1993 Journal: Proc.Natl.Acad.Sci.USA / Year: 1993Title: The Conformation of the Sarcin/Ricin Loop from 28S Ribosomal RNA Authors: Szewczak, A.A. / Moore, P.B. / Chan, Y.L. / Wool, I.G. #2:  Journal: J.Mol.Biol. / Year: 1995 Journal: J.Mol.Biol. / Year: 1995Title: The Sarcin/Ricin Loop, a Modular RNA Authors: Szewczak, A.A. / Moore, P.B. #3:  Journal: Biophys.J. / Year: 1998 Journal: Biophys.J. / Year: 1998Title: Comparison of the Crystal and Solution Structure of Two RNA Oligonucleotides Authors: Rife, J.P. / Stallings, S.C. / Correll, C.C. / Dallas, A. / Steitz, T.A. / Moore, P.B. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  430d.cif.gz 430d.cif.gz | 26.6 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb430d.ent.gz pdb430d.ent.gz | 18 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  430d.json.gz 430d.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/30/430d https://data.pdbj.org/pub/pdb/validation_reports/30/430d ftp://data.pdbj.org/pub/pdb/validation_reports/30/430d ftp://data.pdbj.org/pub/pdb/validation_reports/30/430d | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||||
| Unit cell |
|
- Components
Components
| #1: RNA chain | Mass: 9455.571 Da / Num. of mol.: 1 / Source method: obtained synthetically / Source: (synth.)  | ||
|---|---|---|---|
| #2: Chemical | ChemComp-MG / #3: Water | ChemComp-HOH / | |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.68 Å3/Da / Density % sol: 54.17 % | ||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | pH: 7 / Details: pH 7.0 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Crystal grow | *PLUS Temperature: 20 ℃ / Method: vapor diffusion | ||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  CHESS CHESS  / Beamline: A1 / Beamline: A1 |
| Detector | Type: ADSC / Detector: CCD / Date: Nov 15, 1997 / Details: SILICON 111 BENDING MIRRO |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Relative weight: 1 |
| Reflection | Resolution: 2.1→30 Å / Num. obs: 5854 / % possible obs: 89.6 % / Observed criterion σ(I): -3 / Redundancy: 3.8 % / Biso Wilson estimate: 59.6 Å2 / Rmerge(I) obs: 0.043 / Rsym value: 0.043 / Net I/σ(I): 32.5 |
| Reflection shell | Resolution: 2.1→2.18 Å / Rmerge(I) obs: 0.217 / Mean I/σ(I) obs: 1.9 / Rsym value: 0.217 / % possible all: 88.8 |
| Reflection | *PLUS Num. obs: 6036 / % possible obs: 92.7 % / Num. measured all: 22518 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure: MAD PHASING METHOD / Resolution: 2.1→30 Å / Rfactor Rfree error: 0.013 / Data cutoff high absF: 30991867.71 / Data cutoff low absF: 0 / Cross valid method: THROUGHOUT / σ(F): 0
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 60.3 Å2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine analyze |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.1→30 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 2.1→2.16 Å / Rfactor Rfree error: 0.092 / Total num. of bins used: 12
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Xplor file |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Software | *PLUS Name: CNS / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS Highest resolution: 2.1 Å / Lowest resolution: 30 Å / Rfactor obs: 0.28 / Rfactor Rfree: 0.33 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints | *PLUS
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | *PLUS Highest resolution: 2.1 Å |
 Movie
Movie Controller
Controller








 PDBj
PDBj
































