+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 3wjt | ||||||
|---|---|---|---|---|---|---|---|
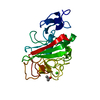
| Title | Crystal structure of the L68D variant of mLolB | ||||||
 Components Components | Outer-membrane lipoprotein LolB | ||||||
 Keywords Keywords | TRANSPORT PROTEIN / LolA/B Fold / Outer membrane | ||||||
| Function / homology | Lipoprotein localisation LolA/LolB/LppX / outer membrane lipoprotein receptor (LolB), chain A / Clam / Mainly Beta / :  Function and homology information Function and homology information | ||||||
| Biological species |  | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.55 Å MOLECULAR REPLACEMENT / Resolution: 1.55 Å | ||||||
 Authors Authors | Takeda, K. / Tokuda, H. / Miki, K. | ||||||
 Citation Citation |  Journal: J.Biol.Chem. / Year: 2014 Journal: J.Biol.Chem. / Year: 2014Title: Roles of the Protruding Loop of Factor B Essential for the Localization of Lipoproteins (LolB) in the Anchoring of Bacterial Triacylated Proteins to the Outer Membran Authors: Hayashi, Y. / Tsurumizu, R. / Tsukahara, J. / Takeda, K. / Narita, S. / Mori, M. / Miki, K. / Tokuda, H. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  3wjt.cif.gz 3wjt.cif.gz | 55.2 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb3wjt.ent.gz pdb3wjt.ent.gz | 38.9 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  3wjt.json.gz 3wjt.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/wj/3wjt https://data.pdbj.org/pub/pdb/validation_reports/wj/3wjt ftp://data.pdbj.org/pub/pdb/validation_reports/wj/3wjt ftp://data.pdbj.org/pub/pdb/validation_reports/wj/3wjt | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  3wjuC  3wjvC  1iwmS C: citing same article ( S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||||||
| Unit cell |
| ||||||||||||
| Components on special symmetry positions |
|
- Components
Components
| #1: Protein | Mass: 20484.809 Da / Num. of mol.: 1 / Fragment: UNP residues 22-207 / Mutation: C1A, L68E Source method: isolated from a genetically manipulated source Source: (gene. exp.)   | ||||
|---|---|---|---|---|---|
| #2: Chemical | | #3: Chemical | ChemComp-CL / | #4: Water | ChemComp-HOH / | |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2 Å3/Da / Density % sol: 38.42 % |
|---|---|
| Crystal grow | Temperature: 277 K / pH: 8 Details: 180mM ammonium sulfate, 25%(w/v) PEG8000, 20mM Tris-HCl, 5.0%(v/v) glycerol, pH 8.0, VAPOR DIFFUSION, SITTING DROP, temperature 277K |
-Data collection
| Diffraction | Mean temperature: 95 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  SPring-8 SPring-8  / Beamline: BL41XU / Wavelength: 1 / Beamline: BL41XU / Wavelength: 1 |
| Detector | Type: ADSC QUANTUM 315 / Detector: CCD / Date: Apr 19, 2008 / Details: MIRRORS |
| Radiation | Monochromator: SI 111 / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1 Å / Relative weight: 1 |
| Reflection | Resolution: 1.55→30 Å / Num. obs: 24057 / % possible obs: 96.1 % / Observed criterion σ(I): 0 / Redundancy: 10.1 % / Biso Wilson estimate: 20.1 Å2 / Rsym value: 0.053 |
| Reflection shell | Resolution: 1.55→1.61 Å / Redundancy: 3.2 % / Rsym value: 0.256 / % possible all: 72.4 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: A CHAIN OF 1IWM Resolution: 1.55→28.17 Å / Rfactor Rfree error: 0.007 / Data cutoff high absF: 133040.49 / Data cutoff low absF: 0 / Isotropic thermal model: RESTRAINED / Cross valid method: THROUGHOUT / σ(F): 0 / Stereochemistry target values: ENGH & HUBER / Details: BULK SOLVENT MODEL USED
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Solvent model: FLAT MODEL / Bsol: 61.4798 Å2 / ksol: 0.4 e/Å3 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 26.2 Å2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine analyze |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.55→28.17 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints NCS | NCS model details: NONE | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 1.55→1.65 Å / Rfactor Rfree error: 0.026 / Total num. of bins used: 6
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Xplor file |
|
 Movie
Movie Controller
Controller











 PDBj
PDBj




