+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 3pdm | ||||||
|---|---|---|---|---|---|---|---|
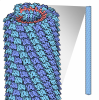
| Title | Hibiscus Latent Singapore virus | ||||||
 Components Components |
| ||||||
 Keywords Keywords | VIRUS / HELICAL VIRUS / VIRUS-RNA COMPLEX | ||||||
| Function / homology |  Function and homology information Function and homology information | ||||||
| Biological species |  Hibiscus latent Singapore virus Hibiscus latent Singapore virus | ||||||
| Method |  FIBER DIFFRACTION / FIBER DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 3.5 Å MOLECULAR REPLACEMENT / Resolution: 3.5 Å | ||||||
 Authors Authors | Tewary, S.K. / Wong, S.M. / Swaminathan, K. | ||||||
 Citation Citation |  Journal: J.Mol.Biol. / Year: 2010 Journal: J.Mol.Biol. / Year: 2010Title: Structure of Hibiscus latent Singapore virus by fiber diffraction: A non-conserved His122 contributes to coat protein stability Authors: Tewary, S.K. / Oda, T. / Kendall, A. / Bian, W. / Stubbs, G. / Wong, S.M. / Swaminathan, K. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  3pdm.cif.gz 3pdm.cif.gz | 52.2 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb3pdm.ent.gz pdb3pdm.ent.gz | 38.4 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  3pdm.json.gz 3pdm.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Summary document |  3pdm_validation.pdf.gz 3pdm_validation.pdf.gz | 362.4 KB | Display |  wwPDB validaton report wwPDB validaton report |
|---|---|---|---|---|
| Full document |  3pdm_full_validation.pdf.gz 3pdm_full_validation.pdf.gz | 398.8 KB | Display | |
| Data in XML |  3pdm_validation.xml.gz 3pdm_validation.xml.gz | 9.8 KB | Display | |
| Data in CIF |  3pdm_validation.cif.gz 3pdm_validation.cif.gz | 13.5 KB | Display | |
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/pd/3pdm https://data.pdbj.org/pub/pdb/validation_reports/pd/3pdm ftp://data.pdbj.org/pub/pdb/validation_reports/pd/3pdm ftp://data.pdbj.org/pub/pdb/validation_reports/pd/3pdm | HTTPS FTP |
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | x 49
| ||||||||
| 2 |
| ||||||||
| 3 | 
| ||||||||
| Unit cell |
| ||||||||
| Symmetry | Helical symmetry: (Circular symmetry: 1 / Dyad axis: no / N subunits divisor: 1 / Num. of operations: 49 / Rise per n subunits: 1.439 Å / Rotation per n subunits: 22.041 °) | ||||||||
| Details | HLSV IS A ROD-SHAPED VIRUS 3000 ANGSTROMS LONG AND 180 ANGSTROMS IN DIAMETER, WITH A CENTRAL HOLE OF DIAMETER 40 ANGSTROMS. APPROXIMATELY 2150 IDENTICAL PROTEIN SUBUNITS OF MOLECULAR WEIGHT 17500 FORM A RIGHT-HANDED HELIX OF PITCH 23.5 ANGSTROMS AND LENGTH 70.5 ANGTROMS WITH 49 SUBUNITS IN THREE TURNS. A SINGLE STRAND OF RNA FOLLOWS THE BASIC HELIX BETWEEN THE PROTEIN SUBUNITS AT A DISTANCE OF 40 ANGSTROMS. THERE ARE THREE NUCLEOTIDES BOUND TO EACH PROTEIN SUBUNIT. THE ASSEMBLY REPRESENTED IN THIS ENTRY HAS REGULAR HELICAL SYMMETRY WITH THE FOLLOWING PARAMETERS: ROTATION PER SUBUNIT (TWIST) = 1080.00/49 DEGREES RISE PER SUBUNIT (HEIGHT) = 70.5/49 ANGSTROMS COORDINATES FOR A COMPLETE MULTIMER REPRESENTING THE KNOWN BIOLOGICALLY SIGNIFICANT OLIGOMERIZATION STATE OF THE MOLECULE CAN BE GENERATED BY APPLYING BIOMT TRANSFORMATIONS GIVEN BELOW. BOTH NON-CRYSTALLOGRAPHIC AND CRYSTALLOGRAPHIC OPERATIONS ARE GIVEN. |
- Components
Components
| #1: RNA chain | Mass: 958.660 Da / Num. of mol.: 1 / Source method: obtained synthetically / Details: synthetic RNA |
|---|---|
| #2: Protein | Mass: 18079.273 Da / Num. of mol.: 1 / Source method: isolated from a natural source / Source: (natural)  Hibiscus latent Singapore virus / References: UniProt: Q8BE68 Hibiscus latent Singapore virus / References: UniProt: Q8BE68 |
-Experimental details
-Experiment
| Experiment | Method:  FIBER DIFFRACTION / Number of used crystals: 1 FIBER DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal grow | Method: oriented sol / Details: Oriented sol |
|---|
-Data collection
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  APS APS  / Beamline: 14-BM-C / Wavelength: 0.9002 Å / Beamline: 14-BM-C / Wavelength: 0.9002 Å |
|---|---|
| Detector | Type: ADSC QUANTUM 315 / Detector: CCD |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.9002 Å / Relative weight: 1 |
| Reflection | Resolution: 3.5→50 Å / Num. all: 3486 / Num. obs: 3486 / % possible obs: 100 % / Observed criterion σ(F): 1 |
- Processing
Processing
| Software |
| |||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT / Resolution: 3.5→50 Å / σ(F): 1 / Stereochemistry target values: Engh & Huber MOLECULAR REPLACEMENT / Resolution: 3.5→50 Å / σ(F): 1 / Stereochemistry target values: Engh & HuberDetails: THE STRUCTURE WAS DETERMINED BY FIBER DIFFRACTION USING MOLECULAR REPLACEMENT WITH LAYER-LINE SPLITTING, SOLVENT FLATTENING REFINEMENT AND RESTRAINED LEAST SQUARES COORDINATE REFINEMENT. THE ...Details: THE STRUCTURE WAS DETERMINED BY FIBER DIFFRACTION USING MOLECULAR REPLACEMENT WITH LAYER-LINE SPLITTING, SOLVENT FLATTENING REFINEMENT AND RESTRAINED LEAST SQUARES COORDINATE REFINEMENT. THE STRUCTURE INCLUDES 162 OF THE 163 AMINO ACIDS AND THREE RNA NUCLEOTIDES MODELLED AS GAA BUT REPRESENTING THE ENTIRE NUCLEIC ACID CONTENT.
| |||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 3.5→50 Å
|
 Movie
Movie Controller
Controller








 PDBj
PDBj
