[English] 日本語
 Yorodumi
Yorodumi- PDB-3obu: Crystal structure of the Tsg101 UEV domain in complex with a HIV-... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 3obu | ||||||
|---|---|---|---|---|---|---|---|
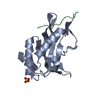
| Title | Crystal structure of the Tsg101 UEV domain in complex with a HIV-1 PTAP peptide | ||||||
 Components Components |
| ||||||
 Keywords Keywords | PROTEIN TRANSPORT / Protein tranport / ubiquitin / HIV-1 Gag | ||||||
| Function / homology |  Function and homology information Function and homology informationpositive regulation of viral budding via host ESCRT complex / positive regulation of ubiquitin-dependent endocytosis / extracellular transport / ESCRT I complex / negative regulation of epidermal growth factor-activated receptor activity / regulation of extracellular exosome assembly / viral budding / regulation of MAP kinase activity / exosomal secretion / protein transport to vacuole involved in ubiquitin-dependent protein catabolic process via the multivesicular body sorting pathway ...positive regulation of viral budding via host ESCRT complex / positive regulation of ubiquitin-dependent endocytosis / extracellular transport / ESCRT I complex / negative regulation of epidermal growth factor-activated receptor activity / regulation of extracellular exosome assembly / viral budding / regulation of MAP kinase activity / exosomal secretion / protein transport to vacuole involved in ubiquitin-dependent protein catabolic process via the multivesicular body sorting pathway / membrane fission / positive regulation of exosomal secretion / multivesicular body assembly / ubiquitin-dependent protein catabolic process via the multivesicular body sorting pathway / Flemming body / virion binding / endosome to lysosome transport / viral budding via host ESCRT complex / negative regulation of epidermal growth factor receptor signaling pathway / viral release from host cell / autophagosome maturation / keratinocyte differentiation / multivesicular body / Endosomal Sorting Complex Required For Transport (ESCRT) / Membrane binding and targetting of GAG proteins / HCMV Late Events / macroautophagy / ubiquitin binding / regulation of cell growth / Late endosomal microautophagy / Budding and maturation of HIV virion / host multivesicular body / calcium-dependent protein binding / transcription corepressor activity / late endosome / late endosome membrane / viral nucleocapsid / early endosome membrane / early endosome / endosome / endosome membrane / regulation of cell cycle / negative regulation of cell population proliferation / viral translational frameshifting / cell division / ubiquitin protein ligase binding / centrosome / protein-containing complex binding / host cell plasma membrane / nucleolus / host cell nucleus / virion membrane / structural molecule activity / negative regulation of transcription by RNA polymerase II / protein homodimerization activity / DNA binding / RNA binding / extracellular exosome / zinc ion binding / plasma membrane / cytoplasm / cytosol Similarity search - Function | ||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.6 Å MOLECULAR REPLACEMENT / Resolution: 1.6 Å | ||||||
 Authors Authors | Im, Y.J. / Hurley, J.H. | ||||||
 Citation Citation |  Journal: Structure / Year: 2010 Journal: Structure / Year: 2010Title: Crystallographic and Functional Analysis of the ESCRT-I /HIV-1 Gag PTAP Interaction. Authors: Im, Y.J. / Kuo, L. / Ren, X. / Burgos, P.V. / Zhao, X.Z. / Liu, F. / Burke, T.R. / Bonifacino, J.S. / Freed, E.O. / Hurley, J.H. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  3obu.cif.gz 3obu.cif.gz | 47.3 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb3obu.ent.gz pdb3obu.ent.gz | 31.6 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  3obu.json.gz 3obu.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/ob/3obu https://data.pdbj.org/pub/pdb/validation_reports/ob/3obu ftp://data.pdbj.org/pub/pdb/validation_reports/ob/3obu ftp://data.pdbj.org/pub/pdb/validation_reports/ob/3obu | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  3obqC  3obsC  3obxC  2f0rS C: citing same article ( S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 16501.195 Da / Num. of mol.: 1 / Fragment: N-terminal UEV domain (UNP residues 2 to 145) / Mutation: 43VFNDGS48 -> GTG Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Gene: TSG101 / Plasmid: pGST2 / Production host: Homo sapiens (human) / Gene: TSG101 / Plasmid: pGST2 / Production host:  |
|---|---|
| #2: Protein/peptide | Mass: 965.997 Da / Num. of mol.: 1 / Fragment: HIV-1 Gag PTAP motif (5-13) / Source method: obtained synthetically Details: This sequence occurs naturally in human immunodeficiency virus type 1. References: UniProt: Q72497 |
| #3: Water | ChemComp-HOH / |
| Sequence details | ENGINEERED |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 1.88 Å3/Da / Density % sol: 34.45 % |
|---|---|
| Crystal grow | Temperature: 298 K / Method: vapor diffusion, hanging drop / pH: 7.5 Details: 0.1M HEPES-NaOH (pH7.5), 25% PEG 3350, 0.2M Sodium Nitrate, VAPOR DIFFUSION, HANGING DROP, temperature 298K |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  APS APS  / Beamline: 22-BM / Wavelength: 1 Å / Beamline: 22-BM / Wavelength: 1 Å |
| Detector | Type: MARMOSAIC 300 mm CCD / Detector: CCD / Date: Aug 7, 2010 / Details: mirrors |
| Radiation | Monochromator: Si 111 CHANNEL / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1 Å / Relative weight: 1 |
| Reflection | Resolution: 1.6→50 Å / Num. all: 18250 / Num. obs: 17689 / % possible obs: 98.8 % / Observed criterion σ(F): 3 / Observed criterion σ(I): 3 / Redundancy: 6.6 % / Biso Wilson estimate: 14.8 Å2 / Rmerge(I) obs: 0.076 / Net I/σ(I): 30.4 |
| Reflection shell | Resolution: 1.6→1.63 Å / Redundancy: 4.9 % / Rmerge(I) obs: 0.328 / Mean I/σ(I) obs: 4.1 / Num. unique all: 799 / % possible all: 93.5 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB ENTRY 2F0R Resolution: 1.6→42.72 Å / Rfactor Rfree error: 0.008 / Data cutoff high absF: 874513.6 / Data cutoff low absF: 0 / Isotropic thermal model: RESTRAINED / Cross valid method: THROUGHOUT / σ(F): 0 / Stereochemistry target values: Engh & Huber
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Solvent model: FLAT MODEL / Bsol: 39.613 Å2 / ksol: 0.358431 e/Å3 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 17.8 Å2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine analyze |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.6→42.72 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints NCS | NCS model details: NONE | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 1.6→1.7 Å / Rfactor Rfree error: 0.024 / Total num. of bins used: 6
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Xplor file |
|
 Movie
Movie Controller
Controller












 PDBj
PDBj






