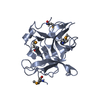
Entry Database : PDB / ID : 3ejfTitle Crystal structure of IBV X-domain at pH 8.5 Non-structural protein 3 Keywords / / / / / / / / Function / homology Function Domain/homology Component
/ / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / Biological species Method / / / Resolution : 1.6 Å Authors Piotrowski, Y. / Hansen, G. / Hilgenfeld, R. Journal : Protein Sci. / Year : 2009Title : Crystal structures of the X-domains of a Group-1 and a Group-3 coronavirus reveal that ADP-ribose-binding may not be a conserved property.Authors : Piotrowski, Y. / Hansen, G. / Boomaars-van der Zanden, A.L. / Snijder, E.J. / Gorbalenya, A.E. / Hilgenfeld, R. History Deposition Sep 18, 2008 Deposition site / Processing site Revision 1.0 Sep 30, 2008 Provider / Type Revision 1.1 Jul 13, 2011 Group Revision 1.2 Oct 25, 2017 Group / Category Revision 1.3 Feb 21, 2024 Group / Database referencesCategory chem_comp_atom / chem_comp_bond ... chem_comp_atom / chem_comp_bond / database_2 / diffrn_source / struct_ref_seq_dif Item _database_2.pdbx_DOI / _database_2.pdbx_database_accession ... _database_2.pdbx_DOI / _database_2.pdbx_database_accession / _diffrn_source.pdbx_synchrotron_site / _struct_ref_seq_dif.details
Show all Show less
 Open data
Open data Basic information
Basic information Components
Components Keywords
Keywords Function and homology information
Function and homology information Avian infectious bronchitis virus
Avian infectious bronchitis virus X-RAY DIFFRACTION /
X-RAY DIFFRACTION /  SYNCHROTRON /
SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.6 Å
MOLECULAR REPLACEMENT / Resolution: 1.6 Å  Authors
Authors Citation
Citation Journal: Protein Sci. / Year: 2009
Journal: Protein Sci. / Year: 2009 Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 3ejf.cif.gz
3ejf.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb3ejf.ent.gz
pdb3ejf.ent.gz PDB format
PDB format 3ejf.json.gz
3ejf.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads https://data.pdbj.org/pub/pdb/validation_reports/ej/3ejf
https://data.pdbj.org/pub/pdb/validation_reports/ej/3ejf ftp://data.pdbj.org/pub/pdb/validation_reports/ej/3ejf
ftp://data.pdbj.org/pub/pdb/validation_reports/ej/3ejf Links
Links Assembly
Assembly
 Components
Components Avian infectious bronchitis virus (strain Beaudette)
Avian infectious bronchitis virus (strain Beaudette)
 X-RAY DIFFRACTION / Number of used crystals: 1
X-RAY DIFFRACTION / Number of used crystals: 1  Sample preparation
Sample preparation SYNCHROTRON / Site:
SYNCHROTRON / Site:  EMBL/DESY, HAMBURG
EMBL/DESY, HAMBURG  / Beamline: X13 / Wavelength: 0.8081
/ Beamline: X13 / Wavelength: 0.8081  Processing
Processing MOLECULAR REPLACEMENT / Resolution: 1.6→24.82 Å / Cor.coef. Fo:Fc: 0.957 / Cor.coef. Fo:Fc free: 0.945 / Occupancy max: 1 / Occupancy min: 0.5 / SU B: 1.37 / SU ML: 0.05 / Cross valid method: THROUGHOUT / σ(F): 0 / ESU R: 0.089 / ESU R Free: 0.09 / Stereochemistry target values: MAXIMUM LIKELIHOOD / Details: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS
MOLECULAR REPLACEMENT / Resolution: 1.6→24.82 Å / Cor.coef. Fo:Fc: 0.957 / Cor.coef. Fo:Fc free: 0.945 / Occupancy max: 1 / Occupancy min: 0.5 / SU B: 1.37 / SU ML: 0.05 / Cross valid method: THROUGHOUT / σ(F): 0 / ESU R: 0.089 / ESU R Free: 0.09 / Stereochemistry target values: MAXIMUM LIKELIHOOD / Details: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS Movie
Movie Controller
Controller














 PDBj
PDBj

