[English] 日本語
 Yorodumi
Yorodumi- PDB-3ejc: Full length Receptor Binding Protein from Lactococcal phage TP901-1 -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 3ejc | ||||||
|---|---|---|---|---|---|---|---|
| Title | Full length Receptor Binding Protein from Lactococcal phage TP901-1 | ||||||
 Components Components | Baseplate protein (BPP) | ||||||
 Keywords Keywords | VIRAL PROTEIN / SUGAR BINDING PROTEIN / Lactococcus lactis / siphoviridae / receptor binding protein / phage TP901-1 / P335 species | ||||||
| Function / homology |  Function and homology information Function and homology informationvirus tail, baseplate / cell adhesion / symbiont entry into host cell / virion attachment to host cell Similarity search - Function | ||||||
| Biological species |  Lactococcus phage TP901-1 (virus) Lactococcus phage TP901-1 (virus) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.85 Å MOLECULAR REPLACEMENT / Resolution: 1.85 Å | ||||||
 Authors Authors | Spinelli, S. / Lichiere, J. / Blangy, S. / Sciara, G. / Cambillau, C. / Campanacci, V. | ||||||
 Citation Citation |  Journal: J Biol Chem / Year: 2010 Journal: J Biol Chem / Year: 2010Title: Structure and molecular assignment of lactococcal phage TP901-1 baseplate. Authors: Cecilia Bebeacua / Patrick Bron / Livia Lai / Christina Skovgaard Vegge / Lone Brøndsted / Silvia Spinelli / Valérie Campanacci / David Veesler / Marin van Heel / Christian Cambillau /  Abstract: P335 lactococcal phages infect the gram(+) bacterium Lactococcus lactis using a large multiprotein complex located at the distal part of the tail and termed baseplate (BP). The BP harbors the ...P335 lactococcal phages infect the gram(+) bacterium Lactococcus lactis using a large multiprotein complex located at the distal part of the tail and termed baseplate (BP). The BP harbors the receptor-binding proteins (RBPs), which allow the specific recognition of saccharidic receptors localized on the host cell surface. We report here the electron microscopic structure of the phage TP901-1 wild-type BP as well as those of two mutants bppL (-) and bppU(-), lacking BppL (the RBPs) or both peripheral BP components (BppL and BppU), respectively. We also achieved an electron microscopic reconstruction of a partial BP complex, formed by BppU and BppL. This complex exhibits a tripod shape and is composed of nine BppLs and three BppUs. These structures, combined with light-scattering measurements, led us to propose that the TP901-1 BP harbors six tripods at its periphery, located around the central tube formed by ORF46 (Dit) hexamers, at its proximal end, and a ORF47 (Tal) trimer at its distal extremity. A total of 54 BppLs (18 RBPs) are thus available to mediate host anchoring with a large apparent avidity. TP901-1 BP exhibits an infection-ready conformation and differs strikingly from the lactococcal phage p2 BP, bearing only 6 RBPs, and which needs a conformational change to reach its activated state. The comparison of several Siphoviridae structures uncovers a close organization of their central BP core whereas striking differences occur at the periphery, leading to diverse mechanisms of host recognition. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  3ejc.cif.gz 3ejc.cif.gz | 46 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb3ejc.ent.gz pdb3ejc.ent.gz | 31.8 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  3ejc.json.gz 3ejc.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/ej/3ejc https://data.pdbj.org/pub/pdb/validation_reports/ej/3ejc ftp://data.pdbj.org/pub/pdb/validation_reports/ej/3ejc ftp://data.pdbj.org/pub/pdb/validation_reports/ej/3ejc | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  1792C  1793C 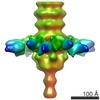 1794C  1795C  2f0cS S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| |||||||||||||||||||||||||||
| Unit cell |
| |||||||||||||||||||||||||||
| Components on special symmetry positions |
|
- Components
Components
| #1: Protein | Mass: 17994.123 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Lactococcus phage TP901-1 (virus) / Gene: bpp / Plasmid: PDEST14 / Production host: Lactococcus phage TP901-1 (virus) / Gene: bpp / Plasmid: PDEST14 / Production host:  |
|---|---|
| #2: Water | ChemComp-HOH / |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.16 Å3/Da / Density % sol: 43.17 % |
|---|---|
| Crystal grow | Temperature: 293 K / Method: vapor diffusion, sitting drop / pH: 6.5 Details: 300 nL of protein at 5 mg/mL were mixed with 100 nL of 20% PEG 8000, 0.2 M Mg Acetate tetrahydrate, 0.1 M Na Cacodylate pH 6.5 using a Cartesian Pixsys Robot, VAPOR DIFFUSION, SITTING DROP, temperature 293K |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  ESRF ESRF  / Beamline: ID14-2 / Wavelength: 0.933 Å / Beamline: ID14-2 / Wavelength: 0.933 Å |
| Detector | Type: ADSC QUANTUM 4 / Detector: CCD / Date: Feb 28, 2008 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.933 Å / Relative weight: 1 |
| Reflection | Resolution: 1.85→30 Å / Num. all: 14330 / Num. obs: 14190 / % possible obs: 99.9 % / Observed criterion σ(F): 0 / Observed criterion σ(I): 0 / Redundancy: 5.5 % / Biso Wilson estimate: 15.8 Å2 / Rmerge(I) obs: 0.103 / Net I/σ(I): 5.6 |
| Reflection shell | Resolution: 1.85→1.95 Å / Redundancy: 5.8 % / Rmerge(I) obs: 0.41 / Mean I/σ(I) obs: 1.7 / Num. unique all: 2025 / % possible all: 99.9 |
- Processing
Processing
| Software |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 2f0c Resolution: 1.85→28.54 Å / Cor.coef. Fo:Fc: 0.943 / Cor.coef. Fo:Fc free: 0.909 / SU B: 5.914 / SU ML: 0.098 / TLS residual ADP flag: LIKELY RESIDUAL / Cross valid method: THROUGHOUT / σ(F): 0 / σ(I): 0 / ESU R: 0.157 / ESU R Free: 0.148 / Stereochemistry target values: MAXIMUM LIKELIHOOD / Details: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Ion probe radii: 0.8 Å / Shrinkage radii: 0.8 Å / VDW probe radii: 1.2 Å / Solvent model: MASK | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 3.853 Å2
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine analyze | Luzzati coordinate error free: 0.148 Å | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.85→28.54 Å
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 1.85→1.898 Å / Total num. of bins used: 20
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS params. | Method: refined / Refine-ID: X-RAY DIFFRACTION
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS group |
|
 Movie
Movie Controller
Controller












 PDBj
PDBj

