[English] 日本語
 Yorodumi
Yorodumi- PDB-2pxl: Variant 9 of Ribonucleoprotein Core of the E. Coli Signal Recogni... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2pxl | ||||||
|---|---|---|---|---|---|---|---|
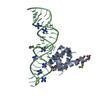
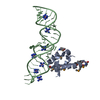
| Title | Variant 9 of Ribonucleoprotein Core of the E. Coli Signal Recognition Particle | ||||||
 Components Components |
| ||||||
 Keywords Keywords | SIGNALING PROTEIN/RNA / GU PAIR / HEXAMINE / RNA PHASING / RNA / CATION BINDING / SIGNALING PROTEIN-RNA COMPLEX | ||||||
| Function / homology |  Function and homology information Function and homology informationsignal recognition particle / signal-recognition-particle GTPase / 7S RNA binding / SRP-dependent cotranslational protein targeting to membrane / protein targeting to membrane / ribonucleoprotein complex / GTPase activity / GTP binding / cytosol Similarity search - Function | ||||||
| Biological species |  | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  MOLECULAR REPLACEMENT / Resolution: 2.5 Å MOLECULAR REPLACEMENT / Resolution: 2.5 Å | ||||||
 Authors Authors | Keel, A.Y. / Rambo, R.P. / Batey, R.T. / Kieft, J.S. | ||||||
 Citation Citation |  Journal: Structure / Year: 2007 Journal: Structure / Year: 2007Title: A General Strategy to Solve the Phase Problem in RNA Crystallography. Authors: Keel, A.Y. / Rambo, R.P. / Batey, R.T. / Kieft, J.S. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2pxl.cif.gz 2pxl.cif.gz | 56.4 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2pxl.ent.gz pdb2pxl.ent.gz | 37.8 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2pxl.json.gz 2pxl.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/px/2pxl https://data.pdbj.org/pub/pdb/validation_reports/px/2pxl ftp://data.pdbj.org/pub/pdb/validation_reports/px/2pxl ftp://data.pdbj.org/pub/pdb/validation_reports/px/2pxl | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  2pxbC  2pxdC  2pxeC  2pxfC  2pxkC  2pxpC  2pxqC  2pxtC  2pxuC  2pxvC C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: RNA chain | Mass: 15235.127 Da / Num. of mol.: 1 / Fragment: DOMAIN IV / Mutation: G131C, C132G, U134G, A175U, G176U, C177G / Source method: obtained synthetically / Details: synthetic | ||
|---|---|---|---|
| #2: Protein | Mass: 12336.646 Da / Num. of mol.: 1 / Fragment: C TERMINAL DOMAIN (RESIDUES 328-432) Source method: isolated from a genetically manipulated source Source: (gene. exp.)   | ||
| #3: Chemical | ChemComp-NCO / Has protein modification | Y | |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.83 Å3/Da / Density % sol: 56.56 % | ||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | Temperature: 293 K / Method: vapor diffusion, sitting drop / pH: 5.6 Details: 20Mm NaOH-MES pH 5.6, 200mM KCl, 13% Isopropanol, 5mM Cobalt Hexamine, VAPOR DIFFUSION, SITTING DROP, temperature 293K | ||||||||||||||||||||||||||||||||||||||||
| Components of the solutions |
|
-Data collection
| Diffraction |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Diffraction source | Source:  ROTATING ANODE / Type: RIGAKU / Wavelength: 1.5418 Å ROTATING ANODE / Type: RIGAKU / Wavelength: 1.5418 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Detector | Type: RIGAKU RAXIS / Detector: IMAGE PLATE / Date: May 24, 2005 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Radiation wavelength | Wavelength: 1.5418 Å / Relative weight: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reflection | Resolution: 2→39.42 Å / Num. obs: 19880 / % possible obs: 95.3 % / Redundancy: 7.5 % / Rmerge(I) obs: 0.115 / Χ2: 0.95 / Net I/σ(I): 9.4 / Scaling rejects: 1127 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reflection shell |
|
-Phasing
| Phasing MR | Cor.coef. Fo:Fc: 0.801 / Packing: 0.407
|
|---|
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT / Resolution: 2.5→50 Å / σ(F): 0 MOLECULAR REPLACEMENT / Resolution: 2.5→50 Å / σ(F): 0
| ||||||||||||||||||||||||||||
| Solvent computation | Bsol: 44.317 Å2 | ||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 60.879 Å2
| ||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.5→50 Å
| ||||||||||||||||||||||||||||
| Xplor file |
|
 Movie
Movie Controller
Controller









 PDBj
PDBj