+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2l2f | ||||||
|---|---|---|---|---|---|---|---|
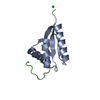
| Title | NMR Structure of GzCVNH (Gibberella zeae CVNH) | ||||||
 Components Components | Cyanovirin-N HOMOLOG | ||||||
 Keywords Keywords | SUGAR BINDING PROTEIN / Cyanovirin-n homolog / lectin / Carbohydrate binding protein | ||||||
| Function / homology | HIV-inactivating Protein, Cyanovirin-n / Cyanovirin-N / Cyanovirin-N / Cyanovirin-N superfamily / CVNH domain / CVNH / Roll / Mainly Beta / Chromosome 3, complete genome Function and homology information Function and homology information | ||||||
| Biological species |  Gibberella zeae (fungus) Gibberella zeae (fungus) | ||||||
| Method | SOLUTION NMR / simulated annealing | ||||||
| Model details | minimized average, model 1 | ||||||
 Authors Authors | Matei, E. / Louis, J.M. / Jee, J.G. / Gronenborn, A.M. | ||||||
 Citation Citation |  Journal: Proteins / Year: 2011 Journal: Proteins / Year: 2011Title: NMR solution structure of a cyanovirin homolog from wheat head blight fungus. Authors: Matei, E. / Louis, J.M. / Jee, J. / Gronenborn, A.M. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2l2f.cif.gz 2l2f.cif.gz | 601.2 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2l2f.ent.gz pdb2l2f.ent.gz | 502.2 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2l2f.json.gz 2l2f.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Summary document |  2l2f_validation.pdf.gz 2l2f_validation.pdf.gz | 343.1 KB | Display |  wwPDB validaton report wwPDB validaton report |
|---|---|---|---|---|
| Full document |  2l2f_full_validation.pdf.gz 2l2f_full_validation.pdf.gz | 482.1 KB | Display | |
| Data in XML |  2l2f_validation.xml.gz 2l2f_validation.xml.gz | 37 KB | Display | |
| Data in CIF |  2l2f_validation.cif.gz 2l2f_validation.cif.gz | 60.8 KB | Display | |
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/l2/2l2f https://data.pdbj.org/pub/pdb/validation_reports/l2/2l2f ftp://data.pdbj.org/pub/pdb/validation_reports/l2/2l2f ftp://data.pdbj.org/pub/pdb/validation_reports/l2/2l2f | HTTPS FTP |
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein | Mass: 11673.413 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Gibberella zeae (fungus) / Production host: Gibberella zeae (fungus) / Production host:  |
|---|
-Experimental details
-Experiment
| Experiment | Method: SOLUTION NMR | ||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
| ||||||||||||||||||||||||||||
| NMR details | Text: THE INITIAL STRUCTURES WERE OBTAINED USING CYANA AUTOMATIC CALCULATION, BASED ON CHEMICAL SHIFT LISTS FROM SEQUENCE-SPECIFIC RESONANCE ASSIGNMENT AND NOES FROM 15N AND 13C-EDITED 3D-NOESY ...Text: THE INITIAL STRUCTURES WERE OBTAINED USING CYANA AUTOMATIC CALCULATION, BASED ON CHEMICAL SHIFT LISTS FROM SEQUENCE-SPECIFIC RESONANCE ASSIGNMENT AND NOES FROM 15N AND 13C-EDITED 3D-NOESY SPECTRA. THROUGHOUT ALL CALCULATIONS,121 BACKBONE TORSION ANGLE CONSTRAINTS DERIVED FROM TALOS, WERE EMPLOYED. CNS WAS USED FOR FURTHER REFINEMENT, USING THE DISTANCE AND DIHEDRAL ANGLE CONSTRAINTS OBTAINED FROM THE FINAL CYCLE OF THE CYANA CALCULATION, AND SEVERAL ADDITIONAL NOE CONSTRAINTS FROM MANUAL CHECKING OF THE 3D NOESY DATA. IN TOTAL, 2401 EXPERIMENTAL NOE-RESTRAINTS (~20 PER RESIDUE) WERE EMPLOYED. |
- Sample preparation
Sample preparation
| Details |
| ||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sample |
| ||||||||||||||||
| Sample conditions | Ionic strength: 20 / pH: 6 / Pressure: AMBIENT / Temperature: 303 K |
-NMR measurement
| NMR spectrometer |
|
|---|
- Processing
Processing
| NMR software |
| ||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method: simulated annealing / Software ordinal: 1 | ||||||||||||||||||||||||||||
| NMR representative | Selection criteria: minimized average | ||||||||||||||||||||||||||||
| NMR ensemble | Conformer selection criteria: structures with the lowest energy Conformers calculated total number: 50 / Conformers submitted total number: 20 |
 Movie
Movie Controller
Controller












 PDBj
PDBj HSQC
HSQC