[English] 日本語
 Yorodumi
Yorodumi- PDB-2l13: mini-haipin of AT basepairs having a C12-alkyl linker forming the... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2l13 | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
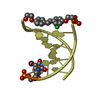
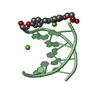
| Title | mini-haipin of AT basepairs having a C12-alkyl linker forming the loop region | ||||||||||||||||||||
 Components Components | DNA (5'-D(* Keywords KeywordsDNA / mini-hairpin / alkyl chain / C12 / dodecyl / synthetic hybrid | Function / homology | DNA / DNA (> 10) |  Function and homology information Function and homology informationMethod | SOLUTION NMR / simulated annealing, matrix relaxation | Model details | minimized average, model 1 |  Authors AuthorsSiegmund, K.H. / Hariharan, M. / Lewis, F.D. |  Citation Citation Journal: J.Phys.Chem.B / Year: 2011 Journal: J.Phys.Chem.B / Year: 2011Title: Conformation of a dodecane DNA hairpin linker. Multiple gauche bonds cover the bases. Authors: Siegmund, K. / Hariharan, M. / Lewis, F.D. History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2l13.cif.gz 2l13.cif.gz | 96.1 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2l13.ent.gz pdb2l13.ent.gz | 77.8 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2l13.json.gz 2l13.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/l1/2l13 https://data.pdbj.org/pub/pdb/validation_reports/l1/2l13 ftp://data.pdbj.org/pub/pdb/validation_reports/l1/2l13 ftp://data.pdbj.org/pub/pdb/validation_reports/l1/2l13 | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: DNA chain | Mass: 3923.739 Da / Num. of mol.: 1 / Source method: obtained synthetically |
|---|
-Experimental details
-Experiment
| Experiment | Method: SOLUTION NMR Details: The C12-alkyl linker enables the short sequence to form a stable mini-hairpin. Ring-currents of the neighboring base-pair made assignment for all methylene protons possible. The position of ...Details: The C12-alkyl linker enables the short sequence to form a stable mini-hairpin. Ring-currents of the neighboring base-pair made assignment for all methylene protons possible. The position of the last terminal base-pair is not well defined due to end-fraying and the resulting lack of a sufficient number of constraints. | ||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
|
- Sample preparation
Sample preparation
| Details |
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Sample |
| |||||||||
| Sample conditions | Ionic strength: 0.1 / pH: 7.2 / Pressure: ambient / Temperature: 293 K |
-NMR measurement
| NMR spectrometer | Type: Varian INOVA / Manufacturer: Varian / Model: INOVA / Field strength: 600 MHz |
|---|
- Processing
Processing
| NMR software |
| ||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method: simulated annealing, matrix relaxation / Software ordinal: 1 | ||||||||||||||||||||||||||||||||
| NMR representative | Selection criteria: minimized average | ||||||||||||||||||||||||||||||||
| NMR ensemble | Conformer selection criteria: structures with the lowest energy Conformers calculated total number: 80 / Conformers submitted total number: 11 |
 Movie
Movie Controller
Controller








 PDBj
PDBj






































 NMRPipe
NMRPipe