| Entry | Database: PDB / ID: 2gpe
|
|---|
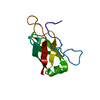
| Title | Structure of the DNA-binding domain of E. Coli Proline Utilization A (PUTA) |
|---|
 Components Components | Bifunctional protein putA |
|---|
 Keywords Keywords | DNA BINDING PROTEIN / PUTA / RIBBON-HELIX-HELIX / DNA-BINDING DOMAIN / PROLINE CATABOLISM / PROLINE UTILIZATION A |
|---|
| Function / homology |  Function and homology information Function and homology information
proline dehydrogenase / proline dehydrogenase activity / L-glutamate gamma-semialdehyde dehydrogenase / L-glutamate gamma-semialdehyde dehydrogenase activity / : / DNA-binding transcription repressor activity / cis-regulatory region sequence-specific DNA binding / cytoplasmic side of plasma membrane / flavin adenine dinucleotide binding / response to oxidative stress ...proline dehydrogenase / proline dehydrogenase activity / L-glutamate gamma-semialdehyde dehydrogenase / L-glutamate gamma-semialdehyde dehydrogenase activity / : / DNA-binding transcription repressor activity / cis-regulatory region sequence-specific DNA binding / cytoplasmic side of plasma membrane / flavin adenine dinucleotide binding / response to oxidative stress / sequence-specific DNA binding / negative regulation of DNA-templated transcription / protein homodimerization activity / DNA binding / identical protein binding / plasma membrane / cytosolSimilarity search - Function : / PutA, RHH domain / Proline dehydrogenase PutA, domain I / Proline utilization A proline dehydrogenase N-terminal domain / Proline utilization A proline dehydrogenase N-terminal domain / Met repressor-like / Delta-1-pyrroline-5-carboxylate dehydrogenase 3 / Proline dehydrogenase PutA, domain II / Proline dehydrogenase PutA, domain I/II / DNA-binding domain of Proline dehydrogenase ...: / PutA, RHH domain / Proline dehydrogenase PutA, domain I / Proline utilization A proline dehydrogenase N-terminal domain / Proline utilization A proline dehydrogenase N-terminal domain / Met repressor-like / Delta-1-pyrroline-5-carboxylate dehydrogenase 3 / Proline dehydrogenase PutA, domain II / Proline dehydrogenase PutA, domain I/II / DNA-binding domain of Proline dehydrogenase / Arc Repressor Mutant / Bifunctional protein PutA / Proline dehydrogenase domain / Proline dehydrogenase / : / Arc-type ribbon-helix-helix / FAD-linked oxidoreductase-like / Ribbon-helix-helix / Aldehyde dehydrogenase, glutamic acid active site / Aldehyde dehydrogenases glutamic acid active site. / Aldehyde dehydrogenase, cysteine active site / Aldehyde dehydrogenases cysteine active site. / Aldehyde dehydrogenase domain / Aldehyde dehydrogenase family / Aldehyde dehydrogenase, C-terminal / Aldehyde dehydrogenase, N-terminal / Aldehyde/histidinol dehydrogenase / Orthogonal Bundle / Mainly AlphaSimilarity search - Domain/homology |
|---|
| Biological species |   Escherichia coli (E. coli) Escherichia coli (E. coli) |
|---|
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.9 Å MOLECULAR REPLACEMENT / Resolution: 1.9 Å |
|---|
 Authors Authors | Jenkins, J.L. / Larson, J. / Tanner, J.J. |
|---|
 Citation Citation |  Journal: Protein Sci. / Year: 2006 Journal: Protein Sci. / Year: 2006
Title: Crystal structures of the DNA-binding domain of Escherichia coli proline utilization A flavoprotein and analysis of the role of Lys9 in DNA recognition.
Authors: Larson, J.D. / Jenkins, J.L. / Schuermann, J.P. / Zhou, Y. / Becker, D.F. / Tanner, J.J. |
|---|
| History | | Deposition | Apr 17, 2006 | Deposition site: RCSB / Processing site: RCSB |
|---|
| Revision 1.0 | Feb 20, 2007 | Provider: repository / Type: Initial release |
|---|
| Revision 1.1 | May 1, 2008 | Group: Version format compliance |
|---|
| Revision 1.2 | Jul 13, 2011 | Group: Advisory / Version format compliance |
|---|
| Revision 1.3 | Aug 30, 2023 | Group: Data collection / Database references ...Data collection / Database references / Derived calculations / Refinement description
Category: chem_comp_atom / chem_comp_bond ...chem_comp_atom / chem_comp_bond / database_2 / pdbx_initial_refinement_model / struct_site
Item: _database_2.pdbx_DOI / _database_2.pdbx_database_accession ..._database_2.pdbx_DOI / _database_2.pdbx_database_accession / _struct_site.pdbx_auth_asym_id / _struct_site.pdbx_auth_comp_id / _struct_site.pdbx_auth_seq_id |
|---|
|
|---|
 Yorodumi
Yorodumi Open data
Open data Basic information
Basic information Components
Components Keywords
Keywords Function and homology information
Function and homology information
 X-RAY DIFFRACTION /
X-RAY DIFFRACTION /  SYNCHROTRON /
SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.9 Å
MOLECULAR REPLACEMENT / Resolution: 1.9 Å  Authors
Authors Citation
Citation Journal: Protein Sci. / Year: 2006
Journal: Protein Sci. / Year: 2006 Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 2gpe.cif.gz
2gpe.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb2gpe.ent.gz
pdb2gpe.ent.gz PDB format
PDB format 2gpe.json.gz
2gpe.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads https://data.pdbj.org/pub/pdb/validation_reports/gp/2gpe
https://data.pdbj.org/pub/pdb/validation_reports/gp/2gpe ftp://data.pdbj.org/pub/pdb/validation_reports/gp/2gpe
ftp://data.pdbj.org/pub/pdb/validation_reports/gp/2gpe
 Links
Links Assembly
Assembly


 Components
Components

 X-RAY DIFFRACTION / Number of used crystals: 1
X-RAY DIFFRACTION / Number of used crystals: 1  Sample preparation
Sample preparation SYNCHROTRON / Site:
SYNCHROTRON / Site:  ALS
ALS  / Beamline: 4.2.2 / Wavelength: 1.2398 Å
/ Beamline: 4.2.2 / Wavelength: 1.2398 Å Processing
Processing MOLECULAR REPLACEMENT
MOLECULAR REPLACEMENT Movie
Movie Controller
Controller











 PDBj
PDBj








