+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2f4u | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Asite RNA + designer antibiotic | ||||||||||||||||||
 Components Components | 5'-R(* Keywords KeywordsRNA / Asite RNA / designer antibiotic | Function / homology | Chem-AB6 / RNA / RNA (> 10) |  Function and homology information Function and homology informationMethod |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  MOLECULAR REPLACEMENT / Resolution: 2.6 Å MOLECULAR REPLACEMENT / Resolution: 2.6 Å  Authors AuthorsMurray, J.B. / Meroueh, S.O. / Russell, R.J. / Lentzen, G. / Haddad, J. / Mobashery, S. |  Citation Citation Journal: Chem.Biol. / Year: 2006 Journal: Chem.Biol. / Year: 2006Title: Interactions of designer antibiotics and the bacterial ribosomal aminoacyl-tRNA site Authors: Murray, J.B. / Meroueh, S.O. / Russell, R.J. / Lentzen, G. / Haddad, J. / Mobashery, S. History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2f4u.cif.gz 2f4u.cif.gz | 34.2 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2f4u.ent.gz pdb2f4u.ent.gz | 23.7 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2f4u.json.gz 2f4u.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/f4/2f4u https://data.pdbj.org/pub/pdb/validation_reports/f4/2f4u ftp://data.pdbj.org/pub/pdb/validation_reports/f4/2f4u ftp://data.pdbj.org/pub/pdb/validation_reports/f4/2f4u | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 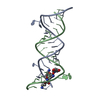
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: RNA chain | Mass: 6743.077 Da / Num. of mol.: 2 / Source method: obtained synthetically / Details: A-site RNA #2: Chemical | ChemComp-AB6 / ( | #3: Water | ChemComp-HOH / | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.55 Å3/Da / Density % sol: 51.79 % |
|---|
-Data collection
| Diffraction | Mean temperature: 100 K | |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Diffraction source | Source:  ROTATING ANODE / Type: RIGAKU RU300 / Wavelength: 1.54 Å ROTATING ANODE / Type: RIGAKU RU300 / Wavelength: 1.54 Å | |||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Detector | Date: Jun 1, 2002 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray | |||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Radiation wavelength | Wavelength: 1.54 Å / Relative weight: 1 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reflection | Resolution: 2.6→23.23 Å / Num. all: 4451 / Num. obs: 4451 / % possible obs: 97.6 % / Observed criterion σ(F): 0 / Observed criterion σ(I): 0 / Rmerge(I) obs: 0.092 / Net I/σ(I): 5.8 / Scaling rejects: 300 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reflection shell |
|
- Processing
Processing
| Software |
| |||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT / Resolution: 2.6→23.23 Å / σ(F): 161 / Stereochemistry target values: Engh & Huber MOLECULAR REPLACEMENT / Resolution: 2.6→23.23 Å / σ(F): 161 / Stereochemistry target values: Engh & Huber
| |||||||||||||||||||||||||
| Solvent computation | Bsol: 95.919 Å2 | |||||||||||||||||||||||||
| Displacement parameters | Biso mean: 56.636 Å2
| |||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.6→23.23 Å
| |||||||||||||||||||||||||
| Xplor file |
|
 Movie
Movie Controller
Controller







 PDBj
PDBj