+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1zim | ||||||
|---|---|---|---|---|---|---|---|
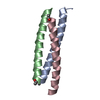
| Title | GCN4-LEUCINE ZIPPER CORE MUTANT ASN16GLN IN THE TRIMERIC STATE | ||||||
 Components Components | GENERAL CONTROL PROTEIN GCN4 | ||||||
 Keywords Keywords | LEUCINE ZIPPER / AMINO-ACID BIOSYNTHESIS / TRANSCRIPTION REGULATION / ACTIVATOR / DNA-BINDING / NUCLEAR PROTEIN / COILED COIL | ||||||
| Function / homology |  Function and homology information Function and homology informationFCERI mediated MAPK activation / protein localization to nuclear periphery / Activation of the AP-1 family of transcription factors / negative regulation of ribosomal protein gene transcription by RNA polymerase II / positive regulation of cellular response to amino acid starvation / response to amino acid starvation / mediator complex binding / Oxidative Stress Induced Senescence / amino acid biosynthetic process / TFIID-class transcription factor complex binding ...FCERI mediated MAPK activation / protein localization to nuclear periphery / Activation of the AP-1 family of transcription factors / negative regulation of ribosomal protein gene transcription by RNA polymerase II / positive regulation of cellular response to amino acid starvation / response to amino acid starvation / mediator complex binding / Oxidative Stress Induced Senescence / amino acid biosynthetic process / TFIID-class transcription factor complex binding / positive regulation of RNA polymerase II transcription preinitiation complex assembly / positive regulation of transcription initiation by RNA polymerase II / cellular response to nutrient levels / cellular response to amino acid starvation / RNA polymerase II transcription regulator complex / DNA-binding transcription activator activity, RNA polymerase II-specific / transcription regulator complex / sequence-specific DNA binding / RNA polymerase II-specific DNA-binding transcription factor binding / DNA-binding transcription factor activity, RNA polymerase II-specific / intracellular signal transduction / RNA polymerase II cis-regulatory region sequence-specific DNA binding / DNA-binding transcription factor activity / chromatin binding / negative regulation of transcription by RNA polymerase II / positive regulation of transcription by RNA polymerase II / identical protein binding / nucleus Similarity search - Function | ||||||
| Biological species |  | ||||||
| Method |  X-RAY DIFFRACTION / Resolution: 2 Å X-RAY DIFFRACTION / Resolution: 2 Å | ||||||
 Authors Authors | Gonzalez Junior, L. / Woolfson, D.N. / Alber, T. | ||||||
 Citation Citation |  Journal: Nat.Struct.Biol. / Year: 1996 Journal: Nat.Struct.Biol. / Year: 1996Title: Buried polar residues and structural specificity in the GCN4 leucine zipper. Authors: Gonzalez Jr., L. / Woolfson, D.N. / Alber, T. #1:  Journal: Nat.Struct.Biol. / Year: 1996 Journal: Nat.Struct.Biol. / Year: 1996Title: An Engineered Allosteric Switch in Leucine-Zipper Oligomerization Authors: Gonzalez Junior, L. / Plecs, J.J. / Alber, T. #2:  Journal: Nature / Year: 1994 Journal: Nature / Year: 1994Title: Crystal Structure of an Isoleucine-Zipper Trimer Authors: Harbury, P.B. / Kim, P.S. / Alber, T. #3:  Journal: Science / Year: 1993 Journal: Science / Year: 1993Title: A Switch between Two-, Three-, and Four-Stranded Coiled Coils in GCN4 Leucine Zipper Mutants Authors: Harbury, P.B. / Zhang, T. / Kim, P.S. / Alber, T. #4:  Journal: Science / Year: 1991 Journal: Science / Year: 1991Title: X-Ray Structure of the GCN4 Leucine Zipper, a Two-Stranded, Parallel Coiled Coil Authors: O'Shea, E.K. / Klemm, J.D. / Kim, P.S. / Alber, T. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1zim.cif.gz 1zim.cif.gz | 31.5 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1zim.ent.gz pdb1zim.ent.gz | 22.3 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1zim.json.gz 1zim.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/zi/1zim https://data.pdbj.org/pub/pdb/validation_reports/zi/1zim ftp://data.pdbj.org/pub/pdb/validation_reports/zi/1zim ftp://data.pdbj.org/pub/pdb/validation_reports/zi/1zim | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein/peptide | Mass: 4045.727 Da / Num. of mol.: 3 / Mutation: N16Q Source method: isolated from a genetically manipulated source Details: THIS STRUCTURE IS IN THE TRIMERIC STATE Source: (gene. exp.)  References: UniProt: P03069 #2: Water | ChemComp-HOH / | Has protein modification | Y | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION X-RAY DIFFRACTION |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 1.77 Å3/Da / Density % sol: 31 % | ||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | *PLUS Temperature: 24 ℃ / Method: vapor diffusion | ||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction source | Source: SEALED TUBE / Wavelength: 1.5418 |
|---|---|
| Detector | Type: RIGAKU RAXIS / Detector: IMAGE PLATE / Date: May 1, 1994 |
| Radiation | Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.5418 Å / Relative weight: 1 |
| Reflection | Num. obs: 5493 / % possible obs: 87 % / Observed criterion σ(I): 2 / Redundancy: 2.2 % / Rmerge(I) obs: 0.077 |
| Reflection | *PLUS Highest resolution: 2 Å / Lowest resolution: 6 Å / Num. measured all: 12314 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Resolution: 2→6 Å / σ(F): 2 /
| ||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2→6 Å
| ||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||
| Refinement | *PLUS Rfactor obs: 0.178 | ||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS |
 Movie
Movie Controller
Controller














 PDBj
PDBj


