[English] 日本語
 Yorodumi
Yorodumi- PDB-1yrn: CRYSTAL STRUCTURE OF THE MATA1/MATALPHA2 HOMEODOMAIN HETERODIMER ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1yrn | ||||||
|---|---|---|---|---|---|---|---|
| Title | CRYSTAL STRUCTURE OF THE MATA1/MATALPHA2 HOMEODOMAIN HETERODIMER BOUND TO DNA | ||||||
 Components Components |
| ||||||
 Keywords Keywords | DNA BINDING PROTEIN/DNA / COMPLEX / DNA-BINDING PROTEIN / DNA / DNA BINDING PROTEIN-DNA COMPLEX | ||||||
| Function / homology |  Function and homology information Function and homology information: / regulation of mating-type specific transcription, DNA-templated / RNA polymerase II transcription repressor complex / DNA binding, bending / DNA-binding transcription repressor activity, RNA polymerase II-specific / sequence-specific DNA binding / DNA-binding transcription factor activity, RNA polymerase II-specific / RNA polymerase II cis-regulatory region sequence-specific DNA binding / regulation of transcription by RNA polymerase II / negative regulation of transcription by RNA polymerase II ...: / regulation of mating-type specific transcription, DNA-templated / RNA polymerase II transcription repressor complex / DNA binding, bending / DNA-binding transcription repressor activity, RNA polymerase II-specific / sequence-specific DNA binding / DNA-binding transcription factor activity, RNA polymerase II-specific / RNA polymerase II cis-regulatory region sequence-specific DNA binding / regulation of transcription by RNA polymerase II / negative regulation of transcription by RNA polymerase II / DNA binding / nucleus Similarity search - Function | ||||||
| Biological species |  | ||||||
| Method |  X-RAY DIFFRACTION / Resolution: 2.5 Å X-RAY DIFFRACTION / Resolution: 2.5 Å | ||||||
 Authors Authors | Li, T. / Stark, M.R. / Johnson, A.D. / Wolberger, C. | ||||||
 Citation Citation |  Journal: Science / Year: 1995 Journal: Science / Year: 1995Title: Crystal structure of the MATa1/MAT alpha 2 homeodomain heterodimer bound to DNA. Authors: Li, T. / Stark, M.R. / Johnson, A.D. / Wolberger, C. #1:  Journal: Proteins / Year: 1995 Journal: Proteins / Year: 1995Title: Crystallization and Preliminary X-Ray Diffraction Studies of an A1/alpha2/DNA Ternary Complex Authors: Li, T. / Stark, M. / Johnson, A.D. / Wolberger, C. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1yrn.cif.gz 1yrn.cif.gz | 63.9 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1yrn.ent.gz pdb1yrn.ent.gz | 43.8 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1yrn.json.gz 1yrn.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/yr/1yrn https://data.pdbj.org/pub/pdb/validation_reports/yr/1yrn ftp://data.pdbj.org/pub/pdb/validation_reports/yr/1yrn ftp://data.pdbj.org/pub/pdb/validation_reports/yr/1yrn | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 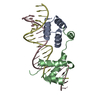
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: DNA chain | Mass: 6395.183 Da / Num. of mol.: 1 / Source method: obtained synthetically |
|---|---|
| #2: DNA chain | Mass: 6484.246 Da / Num. of mol.: 1 / Source method: obtained synthetically |
| #3: Protein | Mass: 7164.493 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Description: T7 PROMOTER / Gene: MAT A1 RESIDUE 66-126 / Plasmid: PMS/K66 / Gene (production host): MAT A1 RESIDUE 66-126 / Production host:  |
| #4: Protein | Mass: 9773.306 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Description: TAC PROMOTER / Gene: MAT ALPHA2 RESIDUE 128-210 / Plasmid: PAV105 / Gene (production host): MAT ALPHA2 RESIDUE 128-210 / Production host:  |
| #5: Water | ChemComp-HOH / |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 3.84 Å3/Da / Density % sol: 67.6 % | ||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal | *PLUS Density % sol: 67.6 % | ||||||||||||||||||||||||||||||||||||||||||
| Crystal grow | *PLUS pH: 6 / Method: vapor diffusion | ||||||||||||||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 94 K |
|---|---|
| Diffraction source | Source:  ROTATING ANODE / Wavelength: 1.5418 Å ROTATING ANODE / Wavelength: 1.5418 Å |
| Detector | Type: RIGAKU RAXIS II / Detector: IMAGE PLATE / Date: Jul 23, 1994 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.5418 Å / Relative weight: 1 |
| Reflection | Resolution: 2.4→20 Å / Num. obs: 18106 / % possible obs: 94 % / Observed criterion σ(I): 1 / Redundancy: 4 % / Rmerge(I) obs: 0.055 |
| Reflection | *PLUS Highest resolution: 2.4 Å / Lowest resolution: 20 Å / % possible obs: 94 % / Redundancy: 4 % / Num. measured all: 76841 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Resolution: 2.5→6 Å / σ(F): 2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 33.65 Å2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine analyze | Luzzati coordinate error obs: 0.3 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.5→6 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Software | *PLUS Name:  X-PLOR / Classification: refinement X-PLOR / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS Highest resolution: 2.5 Å / Lowest resolution: 6 Å / σ(F): 2 / % reflection Rfree: 10 % | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS |
 Movie
Movie Controller
Controller












 PDBj
PDBj







































