[English] 日本語
 Yorodumi
Yorodumi- PDB-1u2u: Nmr solution structure of a designed heterodimeric leucine zipper -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1u2u | ||||||
|---|---|---|---|---|---|---|---|
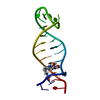
| Title | Nmr solution structure of a designed heterodimeric leucine zipper | ||||||
 Components Components | (General control protein GCN4) x 2 | ||||||
 Keywords Keywords | TRANSCRIPTION / COILED COIL / LEUCINE ZIPPER / INTER-HELICAL ION PAIRING / ELECTROSTATIC INTERACTIONS | ||||||
| Method | SOLUTION NMR / DISTANCE GEOMETRY, REGULARIZATION, SIMULATED ANNEALING, MOLECULAR DYNAMICS SIMULATION IN VACUO | ||||||
 Authors Authors | Marti, D.N. / Bosshard, H.R. | ||||||
 Citation Citation |  Journal: Biochemistry / Year: 2004 Journal: Biochemistry / Year: 2004Title: Inverse electrostatic effect: electrostatic repulsion in the unfolded state stabilizes a leucine zipper. Authors: Marti, D.N. / Bosshard, H.R. #1: Journal: J.Mol.Biol. / Year: 2003 Title: Electrostatic interactions in leucine zippers: thermodynamic analysis of the contributions of Glu and His residues and the effect of mutating salt bridges Authors: Marti, D.N. / Bosshard, H.R. #2:  Journal: Biochemistry / Year: 2000 Journal: Biochemistry / Year: 2000Title: Interhelical ion pairing in coiled coils: solution structure of a heterodimeric leucine zipper and determination of pKa values of Glu side chains Authors: Marti, D.N. / Jelesarov, I. / Bosshard, H.R. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1u2u.cif.gz 1u2u.cif.gz | 517.6 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1u2u.ent.gz pdb1u2u.ent.gz | 437 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1u2u.json.gz 1u2u.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/u2/1u2u https://data.pdbj.org/pub/pdb/validation_reports/u2/1u2u ftp://data.pdbj.org/pub/pdb/validation_reports/u2/1u2u ftp://data.pdbj.org/pub/pdb/validation_reports/u2/1u2u | HTTPS FTP |
|---|
-Related structure data
| Related structure data | |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein/peptide | Mass: 3480.719 Da / Num. of mol.: 1 / Source method: obtained synthetically Details: Synthetic peptide based on the sequence of the leucine zipper domain in GCN4 (baker's yeast) |
|---|---|
| #2: Protein/peptide | Mass: 3508.238 Da / Num. of mol.: 1 / Source method: obtained synthetically Details: synthetic peptide based on the sequence of the leucine zipper domain in GCN4 (baker's yeast) |
| Has protein modification | Y |
-Experimental details
-Experiment
| Experiment | Method: SOLUTION NMR | ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
| ||||||||||||||||||||||||
| NMR details | Text: STRUCTURE WAS DETERMINED USING STANDARD 2D NMR TECHNIQUES |
- Sample preparation
Sample preparation
| Details | Contents: 2.6mM, 90% H2O, 10% D2O / Solvent system: 90% H2O/10% D2O |
|---|---|
| Sample conditions | Ionic strength: 10mM / pH: 5.7 / Pressure: AMBIENT / Temperature: 310.00 K |
-NMR measurement
| NMR spectrometer | Type: Bruker DRX / Manufacturer: Bruker / Model: DRX / Field strength: 600 MHz |
|---|
- Processing
Processing
| NMR software |
| ||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method: DISTANCE GEOMETRY, REGULARIZATION, SIMULATED ANNEALING, MOLECULAR DYNAMICS SIMULATION IN VACUO Software ordinal: 1 Details: STRUCTURES WERE CALCULATED ON THE BASIS OF 1246 NOE DERIVED DISTANCE CONSTRAINTS AND 44 PHI ANGLE RESTRAINTS | ||||||||||||||||||||||||||||
| NMR representative | Selection criteria: closest to the average | ||||||||||||||||||||||||||||
| NMR ensemble | Conformer selection criteria: structures with the least restraint violations, structures with the lowest energy, low deviation of experimental pKa values from pKa derived by continuum electrostatics ...Conformer selection criteria: structures with the least restraint violations, structures with the lowest energy, low deviation of experimental pKa values from pKa derived by continuum electrostatics calculations on structures Conformers calculated total number: 50 / Conformers submitted total number: 27 |
 Movie
Movie Controller
Controller











 PDBj
PDBj HSQC
HSQC