+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1tuh | ||||||
|---|---|---|---|---|---|---|---|
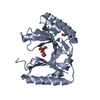
| Title | Structure of Bal32a from a Soil-Derived Mobile Gene Cassette | ||||||
 Components Components | hypothetical protein EGC068 | ||||||
 Keywords Keywords | UNKNOWN FUNCTION / uncultured | ||||||
| Function / homology | SnoaL-like domain / SnoaL-like domain / Nuclear Transport Factor 2; Chain: A, - #50 / NTF2-like domain superfamily / Nuclear Transport Factor 2; Chain: A, / Roll / Alpha Beta / ACETATE ION / SnoaL-like domain-containing protein Function and homology information Function and homology information | ||||||
| Biological species |  uncultured bacterium (environmental samples) uncultured bacterium (environmental samples) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  MIR / Resolution: 1.85 Å MIR / Resolution: 1.85 Å | ||||||
 Authors Authors | Robinson, A. / Wu, P.S.-C. / Harrop, S.J. / Schaeffer, P.M. / Dixon, N.E. / Gillings, M.R. / Holmes, A.J. / Nevalainen, K.M.H. / Otting, G. / Stokes, H.W. ...Robinson, A. / Wu, P.S.-C. / Harrop, S.J. / Schaeffer, P.M. / Dixon, N.E. / Gillings, M.R. / Holmes, A.J. / Nevalainen, K.M.H. / Otting, G. / Stokes, H.W. / Curmi, P.M.G. / Mabbutt, B.C. | ||||||
 Citation Citation |  Journal: J.Mol.Biol. / Year: 2005 Journal: J.Mol.Biol. / Year: 2005Title: Integron-associated Mobile Gene Cassettes Code for Folded Proteins: The Structure of Bal32a, a New Member of the Adaptable alpha+beta Barrel Family Authors: Robinson, A. / Wu, P.S.-C. / Harrop, S.J. / Schaeffer, P.M. / Dosztanyi, Z. / Gillings, M.R. / Holmes, A.J. / Nevalainen, K.M.H. / Stokes, H.W. / Otting, G. / Dixon, N.E. / Curmi, P.M.G. / Mabbutt, B.C. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1tuh.cif.gz 1tuh.cif.gz | 41.5 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1tuh.ent.gz pdb1tuh.ent.gz | 28.6 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1tuh.json.gz 1tuh.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/tu/1tuh https://data.pdbj.org/pub/pdb/validation_reports/tu/1tuh ftp://data.pdbj.org/pub/pdb/validation_reports/tu/1tuh ftp://data.pdbj.org/pub/pdb/validation_reports/tu/1tuh | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| Unit cell |
| ||||||||
| Details | The second part of the biological assembly is generated by the two fold axis: y, x, 1-z. |
- Components
Components
| #1: Protein | Mass: 17767.822 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  uncultured bacterium (environmental samples) uncultured bacterium (environmental samples)Gene: EGC068 / Plasmid: pETMCIII / Production host:  |
|---|---|
| #2: Chemical | ChemComp-ACT / |
| #3: Water | ChemComp-HOH / |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.3 Å3/Da / Density % sol: 47 % |
|---|---|
| Crystal grow | Temperature: 291 K / Method: vapor diffusion, sitting drop / pH: 4.6 Details: ammonium sulfate, sodium acetate, pH 4.6, VAPOR DIFFUSION, SITTING DROP, temperature 291K |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  ROTATING ANODE / Type: ENRAF-NONIUS / Wavelength: 1.5418 Å ROTATING ANODE / Type: ENRAF-NONIUS / Wavelength: 1.5418 Å |
| Detector | Type: MARRESEARCH / Detector: IMAGE PLATE / Date: Jul 25, 2003 / Details: mirrors |
| Radiation | Monochromator: RIGAKU MIRRORS / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.5418 Å / Relative weight: 1 |
| Reflection | Resolution: 1.85→22.5 Å / Num. all: 14962 / Num. obs: 14962 / % possible obs: 100 % / Observed criterion σ(F): 0 / Observed criterion σ(I): 0 |
| Reflection shell | Resolution: 1.85→1.898 Å / % possible all: 100 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MIR / Resolution: 1.85→22.47 Å / Cor.coef. Fo:Fc: 0.96 / Cor.coef. Fo:Fc free: 0.953 / SU B: 2.605 / SU ML: 0.08 / Cross valid method: THROUGHOUT / σ(F): 0 / ESU R: 0.121 / ESU R Free: 0.113 / Stereochemistry target values: MAXIMUM LIKELIHOOD / Details: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS MIR / Resolution: 1.85→22.47 Å / Cor.coef. Fo:Fc: 0.96 / Cor.coef. Fo:Fc free: 0.953 / SU B: 2.605 / SU ML: 0.08 / Cross valid method: THROUGHOUT / σ(F): 0 / ESU R: 0.121 / ESU R Free: 0.113 / Stereochemistry target values: MAXIMUM LIKELIHOOD / Details: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Ion probe radii: 0.8 Å / Shrinkage radii: 0.8 Å / VDW probe radii: 1.2 Å / Solvent model: BABINET MODEL WITH MASK | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 31.676 Å2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.85→22.47 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Highest resolution: 1.85 Å / Num. reflection Rwork: 1034 / Total num. of bins used: 20 |
 Movie
Movie Controller
Controller












 PDBj
PDBj



