+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1sn4 | ||||||
|---|---|---|---|---|---|---|---|
| Title | STRUCTURE OF SCORPION NEUROTOXIN BMK M4 | ||||||
 Components Components | PROTEIN (NEUROTOXIN BMK M4) | ||||||
 Keywords Keywords | TOXIN / NEUROTOXIN / SODIUM CHANNEL INHIBITOR / SCORPION | ||||||
| Function / homology |  Function and homology information Function and homology informationsodium channel inhibitor activity / defense response / toxin activity / extracellular region Similarity search - Function | ||||||
| Biological species |  Mesobuthus martensii (Chinese scorpion) Mesobuthus martensii (Chinese scorpion) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.3 Å MOLECULAR REPLACEMENT / Resolution: 1.3 Å | ||||||
 Authors Authors | He, X.L. / Li, H.M. / Liu, X.Q. / Zeng, Z.H. / Wang, D.C. | ||||||
 Citation Citation |  Journal: J.Mol.Biol. / Year: 1999 Journal: J.Mol.Biol. / Year: 1999Title: Crystal structures of two alpha-like scorpion toxins: non-proline cis peptide bonds and implications for new binding site selectivity on the sodium channel. Authors: He, X.L. / Li, H.M. / Zeng, Z.H. / Liu, X.Q. / Wang, M. / Wang, D.C. #1:  Journal: J.Mol.Biol. / Year: 1994 Journal: J.Mol.Biol. / Year: 1994Title: Crystal Structure of Toxin II from the Scorpion Androctonus Australis Hector Refined at 1.3 A Resolution Authors: Housset, D. / Habersetzer-Rochat, C. / Astier, J.P. / Fontecilla-Camps, J.C. #2:  Journal: Zool.Res.Sinica. / Year: 1989 Journal: Zool.Res.Sinica. / Year: 1989Title: Purification and Partial Characterization of Several New Neurotoxins from East_ Asia Scorpion Crystallographic Studies on an Acidic Toxin from Scorpion Buthus Martensii Karsch Authors: Hu, R.Q. / Wang, M. / LIU, J.N. / LEI, K.J. #3:  Journal: J.Mol.Biol. / Year: 1992 Journal: J.Mol.Biol. / Year: 1992Title: Structure of Scorpion Toxin Variant-3 at 1.2 A Resolution Authors: Zhao, B. / Carson, M. / Ealick, S.E. / Bugg, C.E. #4:  Journal: Toxicon / Year: 1997 Journal: Toxicon / Year: 1997Title: Purification and Sequence Determination of a New Neutral Mammalian Neurotoxin from the Scorpion Buthus Martensii Karsch Authors: Luo, M.J. / Xiong, Y.M. / Wang, M. / Wang, D.C. / Chi, C.W. #5:  Journal: J.Mol.Biol. / Year: 1996 Journal: J.Mol.Biol. / Year: 1996Title: Crystal Structure of an Acidic Neurotoxin from Scorpion Buthus Martensii Karsch at 1.85A Resolution Authors: Li, H.M. / Wang, D.C. / Jin, L. / ZENG, Z.H. / Hu, R.Q. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1sn4.cif.gz 1sn4.cif.gz | 28.5 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1sn4.ent.gz pdb1sn4.ent.gz | 17.8 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1sn4.json.gz 1sn4.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/sn/1sn4 https://data.pdbj.org/pub/pdb/validation_reports/sn/1sn4 ftp://data.pdbj.org/pub/pdb/validation_reports/sn/1sn4 ftp://data.pdbj.org/pub/pdb/validation_reports/sn/1sn4 | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 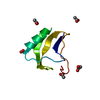
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 7029.957 Da / Num. of mol.: 1 / Source method: isolated from a natural source / Source: (natural)  Mesobuthus martensii (Chinese scorpion) / Organ: TAIL / Secretion: VENOM / References: UniProt: P45698, UniProt: P58328*PLUS Mesobuthus martensii (Chinese scorpion) / Organ: TAIL / Secretion: VENOM / References: UniProt: P45698, UniProt: P58328*PLUS | ||||
|---|---|---|---|---|---|
| #2: Chemical | ChemComp-ACT / #3: Water | ChemComp-HOH / | Has protein modification | Y | |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.07 Å3/Da / Density % sol: 40.6 % | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | pH: 6.8 / Details: pH 6.8 | ||||||||||||||||||||
| Crystal grow | *PLUS Temperature: 23 ℃ / Method: vapor diffusion, hanging drop | ||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 283 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  Photon Factory Photon Factory  / Beamline: BL-6A / Wavelength: 1 / Beamline: BL-6A / Wavelength: 1 |
| Detector | Detector: IMAGE PLATE / Date: Dec 14, 1991 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1 Å / Relative weight: 1 |
| Reflection | Resolution: 1.3→21.3 Å / Num. obs: 10577 / % possible obs: 73.5 % / Redundancy: 2.2 % / Biso Wilson estimate: 20.2 Å2 / Rmerge(I) obs: 0.062 / Net I/σ(I): 3.7 |
| Reflection shell | Resolution: 1.3→1.35 Å / Redundancy: 1.8 % / Rmerge(I) obs: 0.32 / Mean I/σ(I) obs: 2.2 / % possible all: 51.2 |
| Reflection | *PLUS Num. obs: 10578 / Num. measured all: 23268 |
| Reflection shell | *PLUS % possible obs: 60.2 % |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: BMK M1 Resolution: 1.3→20 Å / Data cutoff high absF: 100000 / Data cutoff low absF: 0.1 / Isotropic thermal model: RESTRAINED / Cross valid method: THROUGHOUT / σ(F): 2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 26.9 Å2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.3→20 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 1.3→1.35 Å / Total num. of bins used: 10
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Xplor file |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Software | *PLUS Name:  X-PLOR / Classification: refinement X-PLOR / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS Highest resolution: 1.3 Å / Lowest resolution: 20 Å / σ(F): 2 / % reflection Rfree: 10 % | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS Biso mean: 26.9 Å2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints | *PLUS
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | *PLUS Rfactor Rfree: 0.343 / % reflection Rfree: 10.8 % / Rfactor Rwork: 0.299 |
 Movie
Movie Controller
Controller












 PDBj
PDBj




