[English] 日本語
 Yorodumi
Yorodumi- PDB-1s1k: INFLUENCE OF GROOVE INTERACTIONS ON DNA HOLLIDAY JUNCTION FORMATION -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1s1k | |||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | INFLUENCE OF GROOVE INTERACTIONS ON DNA HOLLIDAY JUNCTION FORMATION | |||||||||||||||||||||||
 Components Components | 5'-D(* Keywords KeywordsDNA / B-DNA / DOUBLE HELIX / 2 / 6-Diaminopurine / Minor groove interactions | Function / homology | DNA |  Function and homology information Function and homology informationMethod |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.9 Å MOLECULAR REPLACEMENT / Resolution: 1.9 Å  Authors AuthorsHays, F.A. / Watson, J. / Ho, P.S. |  Citation Citation Journal: Biochemistry / Year: 2004 Journal: Biochemistry / Year: 2004Title: Influence of minor groove substituents on the structure of DNA holliday junctions. Authors: Hays, F.A. / Jones, Z.J. / Ho, P.S. History |
Remark 300 | BIOMOLECULE: 1 THIS ENTRY CONTAINS THE CRYSTALLOGRAPHIC ASYMMETRIC UNIT WHICH CONSISTS OF 1 CHAIN(S) ...BIOMOLECULE: 1 THIS ENTRY CONTAINS THE CRYSTALLOGRAPHIC ASYMMETRIC UNIT WHICH CONSISTS OF 1 CHAIN(S). THE FULL BIOLOGICAL UNIT CONSISTS OF TWO STRANDS FORMING A B-DNA DUPLEX STRUCTURE. SEE REMARK 350 FOR INFORMATION ON GENERATING THE BIOLOGICAL MOLECULE(S). | |
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1s1k.cif.gz 1s1k.cif.gz | 20.8 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1s1k.ent.gz pdb1s1k.ent.gz | 11.6 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1s1k.json.gz 1s1k.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/s1/1s1k https://data.pdbj.org/pub/pdb/validation_reports/s1/1s1k ftp://data.pdbj.org/pub/pdb/validation_reports/s1/1s1k ftp://data.pdbj.org/pub/pdb/validation_reports/s1/1s1k | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  1s1lC  1p4zS S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 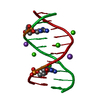
| ||||||||||||
| Unit cell |
| ||||||||||||
| Components on special symmetry positions |
|
- Components
Components
| #1: DNA chain | Mass: 3060.019 Da / Num. of mol.: 1 / Source method: obtained synthetically | ||||
|---|---|---|---|---|---|
| #2: Chemical | | #3: Chemical | ChemComp-NA / | #4: Water | ChemComp-HOH / | |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.02 Å3/Da / Density % sol: 38.5 % | ||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | Method: vapor diffusion, sitting drop / pH: 7.5 Details: 5mM TRIS(HCl), 23mM CaAcetate, 16% MPD, .6mM DNA against 28% MPD at RT, pH 7.50, VAPOR DIFFUSION, SITTING DROP | ||||||||||||||||||||||||||||||||
| Components of the solutions |
|
-Data collection
| Diffraction | Mean temperature: 103 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  APS APS  / Beamline: 14-BM-C / Wavelength: 0.9 Å / Beamline: 14-BM-C / Wavelength: 0.9 Å |
| Detector | Type: ADSC QUANTUM 4 / Detector: CCD / Date: Jan 8, 2002 / Details: BENT CONICAL SI-MIRROR (RH COATING) |
| Radiation | Monochromator: BENT GE(111) MONOCHROMATOR / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.9 Å / Relative weight: 1 |
| Reflection | Resolution: 1.85→17.5 Å / Num. all: 2528 / Num. obs: 2528 / % possible obs: 96.3 % / Observed criterion σ(I): 0 / Biso Wilson estimate: 8.8 Å2 / Rmerge(I) obs: 0.056 |
| Reflection shell | Resolution: 1.85→1.92 Å / Rmerge(I) obs: 0.245 / Mean I/σ(I) obs: 4.98 / % possible all: 72.2 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 1P4Z Resolution: 1.9→17.49 Å / Rfactor Rfree error: 0.017 / Data cutoff high absF: 29235.77 / Data cutoff high rms absF: 29235.77 / Data cutoff low absF: 0 / Isotropic thermal model: RESTRAINED / Cross valid method: THROUGHOUT / σ(F): 0 / σ(I): 0 / Stereochemistry target values: CNS
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 14 Å2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine analyze |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.9→17.49 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 1.9→2.02 Å / Rfactor Rfree error: 0.069 / Total num. of bins used: 6
|
 Movie
Movie Controller
Controller






 PDBj
PDBj






