[English] 日本語
 Yorodumi
Yorodumi- PDB-1qw4: Crystal Structure of Murine Inducible Nitric Oxide Synthase Oxyge... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1qw4 | ||||||
|---|---|---|---|---|---|---|---|
| Title | Crystal Structure of Murine Inducible Nitric Oxide Synthase Oxygenase Domain in complex with N-omega-propyl-L-arginine. | ||||||
 Components Components | Nitric oxide synthase, inducible | ||||||
 Keywords Keywords | OXIDOREDUCTASE / iNOSoxy inhibitor complex | ||||||
| Function / homology |  Function and homology information Function and homology informationNitric oxide stimulates guanylate cyclase / ROS and RNS production in phagocytes / peptidyl-cysteine S-nitrosylation / G protein-coupled receptor signaling pathway coupled to cGMP nucleotide second messenger / cAMP-dependent protein kinase regulator activity / Peroxisomal protein import / prostaglandin secretion / tetrahydrobiopterin binding / : / arginine binding ...Nitric oxide stimulates guanylate cyclase / ROS and RNS production in phagocytes / peptidyl-cysteine S-nitrosylation / G protein-coupled receptor signaling pathway coupled to cGMP nucleotide second messenger / cAMP-dependent protein kinase regulator activity / Peroxisomal protein import / prostaglandin secretion / tetrahydrobiopterin binding / : / arginine binding / superoxide metabolic process / regulation of cytokine production involved in inflammatory response / Fc-gamma receptor signaling pathway involved in phagocytosis / cellular response to cytokine stimulus / regulation of insulin secretion / cortical cytoskeleton / nitric-oxide synthase binding / response to tumor necrosis factor / nitric-oxide synthase (NADPH) / : / blood vessel remodeling / nitric-oxide synthase activity / L-arginine catabolic process / nitric oxide biosynthetic process / negative regulation of blood pressure / response to hormone / positive regulation of interleukin-8 production / response to bacterium / circadian rhythm / Hsp90 protein binding / negative regulation of protein catabolic process / positive regulation of interleukin-6 production / beta-catenin binding / cellular response to type II interferon / regulation of blood pressure / cellular response to xenobiotic stimulus / NADP binding / FMN binding / flavin adenine dinucleotide binding / peroxisome / regulation of cell population proliferation / cellular response to lipopolysaccharide / actin binding / response to lipopolysaccharide / response to hypoxia / calmodulin binding / defense response to bacterium / intracellular signal transduction / cadherin binding / positive regulation of apoptotic process / inflammatory response / negative regulation of gene expression / heme binding / protein kinase binding / perinuclear region of cytoplasm / protein homodimerization activity / : / metal ion binding / identical protein binding / nucleus / plasma membrane / cytoplasm / cytosol Similarity search - Function | ||||||
| Biological species |  | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2.4 Å MOLECULAR REPLACEMENT / Resolution: 2.4 Å | ||||||
 Authors Authors | Fedorov, R. / Hartmann, E. / Ghosh, D.K. / Schlichting, I. | ||||||
 Citation Citation |  Journal: J.Biol.Chem. / Year: 2003 Journal: J.Biol.Chem. / Year: 2003Title: Structural basis for the specificity of the nitric-oxide synthase inhibitors W1400 and Nomega-propyl-L-Arg for the inducible and neuronal isoforms. Authors: Fedorov, R. / Hartmann, E. / Ghosh, D.K. / Schlichting, I. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1qw4.cif.gz 1qw4.cif.gz | 193.9 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1qw4.ent.gz pdb1qw4.ent.gz | 154.2 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1qw4.json.gz 1qw4.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/qw/1qw4 https://data.pdbj.org/pub/pdb/validation_reports/qw/1qw4 ftp://data.pdbj.org/pub/pdb/validation_reports/qw/1qw4 ftp://data.pdbj.org/pub/pdb/validation_reports/qw/1qw4 | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||||||||||||||||||
| 2 | 
| ||||||||||||||||||||||||
| Unit cell |
| ||||||||||||||||||||||||
| Components on special symmetry positions |
|
- Components
Components
-Protein , 1 types, 2 molecules AB
| #1: Protein | Mass: 48498.168 Da / Num. of mol.: 2 Fragment: inducible nitric oxide synthase oxygenase domain (residues 77-495) Source method: isolated from a genetically manipulated source Source: (gene. exp.)   |
|---|
-Non-polymers , 5 types, 361 molecules 


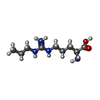





| #2: Chemical | | #3: Chemical | #4: Chemical | #5: Chemical | #6: Water | ChemComp-HOH / | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 4.02 Å3/Da / Density % sol: 69.43 % | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | Temperature: 277 K / Method: vapor diffusion, hanging drop / pH: 7 Details: MES, sodium malonate, EPPS, NaCl, glycerol, H4B, β-mercaptoethanol, N-omega-propyl-L-Arg., pH 7.0, VAPOR DIFFUSION, HANGING DROP, temperature 277K | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Crystal grow | *PLUS Temperature: 4 ℃ / pH: 7.6 / Method: vapor diffusion, hanging drop | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  ESRF ESRF  / Beamline: ID14-1 / Wavelength: 0.934 Å / Beamline: ID14-1 / Wavelength: 0.934 Å |
| Detector | Type: ADSC QUANTUM 4 / Detector: CCD / Date: Oct 2, 2001 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.934 Å / Relative weight: 1 |
| Reflection | Resolution: 2.3→20 Å / Num. obs: 69234 / % possible obs: 95.4 % / Observed criterion σ(F): 2 / Observed criterion σ(I): 2 / Redundancy: 2.7 % / Biso Wilson estimate: 51.3 Å2 / Rsym value: 0.059 / Net I/σ(I): 9.8 |
| Reflection shell | Resolution: 2.3→2.35 Å / Mean I/σ(I) obs: 2.6 / Rsym value: 0.384 / % possible all: 89.4 |
| Reflection | *PLUS Num. obs: 128211 / Num. measured all: 350863 / Rmerge(I) obs: 0.059 |
| Reflection shell | *PLUS % possible obs: 89.4 % / Rmerge(I) obs: 0.384 |
- Processing
Processing
| Software |
| ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT / Resolution: 2.4→8 Å / σ(F): 0 / Stereochemistry target values: Engh & Huber MOLECULAR REPLACEMENT / Resolution: 2.4→8 Å / σ(F): 0 / Stereochemistry target values: Engh & Huber
| ||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.4→8 Å
| ||||||||||||||||||||
| Refinement | *PLUS % reflection Rfree: 5 % | ||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||
| Displacement parameters | *PLUS | ||||||||||||||||||||
| Refine LS restraints | *PLUS
|
 Movie
Movie Controller
Controller


























 PDBj
PDBj






