[English] 日本語
 Yorodumi
Yorodumi- PDB-1qda: Crystal structure of epidoxorubicin-formaldehyde virtual crosslin... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1qda | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
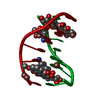
| Title | Crystal structure of epidoxorubicin-formaldehyde virtual crosslink of DNA | ||||||||||||||||||
 Components Components | 5' -D(CP* Keywords KeywordsDNA / DOUBLE HELIX / DRUG-DNA COMPLEX | Function / homology | 4'-EPIDOXORUBICIN / DNA |  Function and homology information Function and homology informationMethod |  X-RAY DIFFRACTION / Resolution: 1.6 Å X-RAY DIFFRACTION / Resolution: 1.6 Å  Authors AuthorsPodell, E.R. / Harrington, D.J. / Taatjes, D.J. / Koch, T.H. |  Citation Citation Journal: Acta Crystallogr.,Sect.D / Year: 1999 Journal: Acta Crystallogr.,Sect.D / Year: 1999Title: Crystal structure of epidoxorubicin-formaldehyde virtual crosslink of DNA and evidence for its formation in human breast-cancer cells. Authors: Podell, E.R. / Harrington, D.J. / Taatjes, D.J. / Koch, T.H. #1: Journal: Curr.Pharm.Des. / Year: 1998 Title: A redox pathway leading to the alkylation of nucleic acids by doxorubicin and related anthracyclines: application to the design of antitumor drugs for resistant cancer. Authors: Taatjes, D.J. / Fenick, D.J. / Gaudiano, G. / Koch, T.H. #2:  Journal: J.Med.Chem. / Year: 1998 Journal: J.Med.Chem. / Year: 1998Title: Epidoxoform: A Hydrolytically More Stable Anthracycline-Formaldehyde Conjugate Toxic to Resistant Tumor Cells Authors: Taatjes, D.J. / Fenick, D.J. / Koch, T.H. History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1qda.cif.gz 1qda.cif.gz | 15.1 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1qda.ent.gz pdb1qda.ent.gz | 9.2 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1qda.json.gz 1qda.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Summary document |  1qda_validation.pdf.gz 1qda_validation.pdf.gz | 743.5 KB | Display |  wwPDB validaton report wwPDB validaton report |
|---|---|---|---|---|
| Full document |  1qda_full_validation.pdf.gz 1qda_full_validation.pdf.gz | 744.9 KB | Display | |
| Data in XML |  1qda_validation.xml.gz 1qda_validation.xml.gz | 3.5 KB | Display | |
| Data in CIF |  1qda_validation.cif.gz 1qda_validation.cif.gz | 4 KB | Display | |
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/qd/1qda https://data.pdbj.org/pub/pdb/validation_reports/qd/1qda ftp://data.pdbj.org/pub/pdb/validation_reports/qd/1qda ftp://data.pdbj.org/pub/pdb/validation_reports/qd/1qda | HTTPS FTP |
-Related structure data
| Related structure data | |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||||
| Unit cell |
|
- Components
Components
| #1: DNA chain | Mass: 1824.232 Da / Num. of mol.: 1 / Source method: obtained synthetically Details: DOXORUBICIN COVALENTLY ATTACHED TO N2(G4) VIA METHYLENE BRIDGE |
|---|---|
| #2: Chemical | ChemComp-DM6 / |
| #3: Water | ChemComp-HOH / |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.72 Å3/Da / Density % sol: 54.82 % | ||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | Temperature: 296 K / Method: vapor diffusion, hanging drop / pH: 6 Details: MPD, FORMALDEHYDE, CACODYLATE, SPERMINE, BARIUM CHLORIDE, pH 6.0, VAPOR DIFFUSION, HANGING DROP, temperature 296K | ||||||||||||||||||||||||||||||||||||||||||||||||
| Components of the solutions |
| ||||||||||||||||||||||||||||||||||||||||||||||||
| Crystal grow | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 298 K |
|---|---|
| Diffraction source | Source:  ROTATING ANODE / Type: RIGAKU RU200 / Wavelength: 1.5418 ROTATING ANODE / Type: RIGAKU RU200 / Wavelength: 1.5418 |
| Detector | Type: RIGAKU RAXIS II / Detector: IMAGE PLATE / Date: Jan 23, 1998 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.5418 Å / Relative weight: 1 |
| Reflection | Resolution: 1.6→50 Å / Num. all: 3101 / Num. obs: 3101 / % possible obs: 92 % / Observed criterion σ(F): 0 / Observed criterion σ(I): 0 / Redundancy: 12 % / Biso Wilson estimate: 17.8 Å2 / Rmerge(I) obs: 0.099 / Net I/σ(I): 6.8 |
| Reflection shell | Resolution: 1.6→1.66 Å / Redundancy: 9.5 % / Rmerge(I) obs: 0.309 / % possible all: 82.7 |
| Reflection | *PLUS Num. measured all: 47174 |
| Reflection shell | *PLUS % possible obs: 82.7 % / Num. unique obs: 2482 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Resolution: 1.6→30 Å / Cross valid method: THROUGHOUT / σ(F): 2 / σ(I): 2 Stereochemistry target values: G. PARKINSON, J. VOJTECHOVSKY, L. CLOWNEY, A.T. BRUNGER, H.M. BERMAN Details: USED ALL THE REFLECTIONS IN THE FINAL SERIES OF REFINEMENT CYCLES
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.6→30 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Software | *PLUS Name: CNS / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS Highest resolution: 1.6 Å / Lowest resolution: 30 Å / σ(F): 2 / % reflection Rfree: 17.3 % / Rfactor all: 0.2164 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS Biso mean: 19.37 Å2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints | *PLUS
|
 Movie
Movie Controller
Controller







 PDBj
PDBj