+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1q1u | ||||||
|---|---|---|---|---|---|---|---|
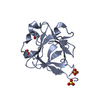
| Title | Crystal structure of human FHF1b (FGF12b) | ||||||
 Components Components | fibroblast growth factor homologous factor 1 | ||||||
 Keywords Keywords | HORMONE/GROWTH FACTOR / FGF-12 / Human / crystal structure / HORMONE-GROWTH FACTOR COMPLEX | ||||||
| Function / homology |  Function and homology information Function and homology informationregulation of voltage-gated sodium channel activity / regulation of neuronal action potential / positive regulation of sodium ion transport / regulation of sodium ion transmembrane transport / cardiac muscle cell action potential involved in contraction / neuromuscular process / Phase 0 - rapid depolarisation / sodium channel regulator activity / JNK cascade / neurogenesis ...regulation of voltage-gated sodium channel activity / regulation of neuronal action potential / positive regulation of sodium ion transport / regulation of sodium ion transmembrane transport / cardiac muscle cell action potential involved in contraction / neuromuscular process / Phase 0 - rapid depolarisation / sodium channel regulator activity / JNK cascade / neurogenesis / adult locomotory behavior / growth factor activity / nervous system development / cell-cell signaling / heparin binding / heart development / chemical synaptic transmission / transmembrane transporter binding / synapse / signal transduction / extracellular space / nucleus / cytoplasm Similarity search - Function | ||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.7 Å MOLECULAR REPLACEMENT / Resolution: 1.7 Å | ||||||
 Authors Authors | Olsen, S.K. / Garbi, M. / Zampieri, N. / Eliseenkova, A.V. / Ornitz, D.M. / Goldfarb, M. / Mohammadi, M. | ||||||
 Citation Citation |  Journal: J.Biol.Chem. / Year: 2003 Journal: J.Biol.Chem. / Year: 2003Title: Fibroblast growth factor (FGF) homologous factors share structural but not functional homology with FGFs Authors: Olsen, S.K. / Garbi, M. / Zampieri, N. / Eliseenkova, A.V. / Ornitz, D.M. / Goldfarb, M. / Mohammadi, M. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1q1u.cif.gz 1q1u.cif.gz | 41 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1q1u.ent.gz pdb1q1u.ent.gz | 27.7 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1q1u.json.gz 1q1u.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Summary document |  1q1u_validation.pdf.gz 1q1u_validation.pdf.gz | 376.9 KB | Display |  wwPDB validaton report wwPDB validaton report |
|---|---|---|---|---|
| Full document |  1q1u_full_validation.pdf.gz 1q1u_full_validation.pdf.gz | 377.8 KB | Display | |
| Data in XML |  1q1u_validation.xml.gz 1q1u_validation.xml.gz | 4.2 KB | Display | |
| Data in CIF |  1q1u_validation.cif.gz 1q1u_validation.cif.gz | 6 KB | Display | |
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/q1/1q1u https://data.pdbj.org/pub/pdb/validation_reports/q1/1q1u ftp://data.pdbj.org/pub/pdb/validation_reports/q1/1q1u ftp://data.pdbj.org/pub/pdb/validation_reports/q1/1q1u | HTTPS FTP |
-Related structure data
| Related structure data |  1ihkS S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 16365.646 Da / Num. of mol.: 1 / Fragment: residues 1-144 / Mutation: C144A Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Description: T7 promoter driven expression / Plasmid: pET / Species (production host): Escherichia coli / Production host: Homo sapiens (human) / Description: T7 promoter driven expression / Plasmid: pET / Species (production host): Escherichia coli / Production host:  | ||
|---|---|---|---|
| #2: Chemical | ChemComp-SO4 / #3: Water | ChemComp-HOH / | |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 1.8 Å3/Da / Density % sol: 31.7 % | ||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | Temperature: 293 K / Method: vapor diffusion, hanging drop / pH: 7.5 Details: 20% PEG 400, 200mM ammonium sulfate, pH 7.5, VAPOR DIFFUSION, HANGING DROP, temperature 293K | ||||||||||||||||||||||||||||||||||||||||||
| Crystal grow | *PLUS Temperature: 20 ℃ / Method: vapor diffusion, hanging drop | ||||||||||||||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 90 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  NSLS NSLS  / Beamline: X4A / Wavelength: 0.920218 / Beamline: X4A / Wavelength: 0.920218 |
| Detector | Type: ADSC QUANTUM 4 / Detector: CCD / Date: Oct 30, 2002 |
| Radiation | Monochromator: Kohzu double crystal monochromator / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.920218 Å / Relative weight: 1 |
| Reflection | Resolution: 1.7→30 Å / Num. all: 13559 / Num. obs: 13171 / % possible obs: 97 % / Observed criterion σ(I): 0 / Rsym value: 0.058 / Net I/σ(I): 7.03 |
| Reflection shell | Resolution: 1.7→1.76 Å / % possible obs: 85 % / % possible all: 85 |
| Reflection | *PLUS Highest resolution: 1.7 Å / Num. measured all: 93909 / Rmerge(I) obs: 0.058 |
| Reflection shell | *PLUS Highest resolution: 1.7 Å / Rmerge(I) obs: 0.365 |
- Processing
Processing
| Software |
| |||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB ID 1IHK Resolution: 1.7→25 Å / Cross valid method: THROUGHOUT / σ(F): 0 / Stereochemistry target values: Engh & Huber
| |||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.7→25 Å
| |||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||
| Refinement | *PLUS Highest resolution: 1.7 Å / % reflection Rfree: 10 % / Rfactor Rfree: 0.24 | |||||||||||||||||||||||||
| Solvent computation | *PLUS | |||||||||||||||||||||||||
| Displacement parameters | *PLUS |
 Movie
Movie Controller
Controller












 PDBj
PDBj








