[English] 日本語
 Yorodumi
Yorodumi- PDB-1ppp: CRYSTAL STRUCTURE OF PAPAIN-E64-C COMPLEX. BINDING DIVERSITY OF E... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1ppp | ||||||
|---|---|---|---|---|---|---|---|
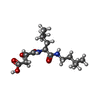
| Title | CRYSTAL STRUCTURE OF PAPAIN-E64-C COMPLEX. BINDING DIVERSITY OF E64-C TO PAPAIN S2 AND S3 SUBSITES | ||||||
 Components Components | PAPAIN | ||||||
 Keywords Keywords | HYDROLASE(SULFHYDRYL PROTEINASE) | ||||||
| Function / homology |  Function and homology information Function and homology informationpapain / serpin family protein binding / cysteine-type peptidase activity / proteolysis Similarity search - Function | ||||||
| Biological species |  | ||||||
| Method |  X-RAY DIFFRACTION / Resolution: 1.9 Å X-RAY DIFFRACTION / Resolution: 1.9 Å | ||||||
 Authors Authors | Ishida, T. | ||||||
 Citation Citation |  Journal: Biochem.J. / Year: 1992 Journal: Biochem.J. / Year: 1992Title: Crystal structure of papain-E64-c complex. Binding diversity of E64-c to papain S2 and S3 subsites. Authors: Kim, M.J. / Yamamoto, D. / Matsumoto, K. / Inoue, M. / Ishida, T. / Mizuno, H. / Sumiya, S. / Kitamura, K. #1:  Journal: J.Biol.Chem. / Year: 1991 Journal: J.Biol.Chem. / Year: 1991Title: Refined X-Ray Structure of Papain(Dot)E-64-C Complex at 2.1-Angstroms Resolution Authors: Yamamoto, D. / Matsumoto, K. / Ohishi, H. / Ishida, T. / Inoue, M. / Kitamura, K. / Mizuno, H. #2:  Journal: FEBS Lett. / Year: 1989 Journal: FEBS Lett. / Year: 1989Title: Mode of Binding of E-64-C, a Potent Thiol Protease Inhibitor, to Papain as Determined by X-Ray Crystal Analysis of the Complex Authors: Matsumoto, K. / Yamamoto, D. / Ohishi, H. / Tomoo, K. / Ishida, T. / Inoue, M. / Sadatome, T. / Kitamura, K. / Mizuno, H. #3:  Journal: FEBS Lett. / Year: 1990 Journal: FEBS Lett. / Year: 1990Title: The Importance of Val-157 Hydrophobic Interaction for Papain Inhibitory Activity of an Epoxysuccinyl Amino Acid Derivative; a Structure-Activity Relationship Based on the Crystal Structure of ...Title: The Importance of Val-157 Hydrophobic Interaction for Papain Inhibitory Activity of an Epoxysuccinyl Amino Acid Derivative; a Structure-Activity Relationship Based on the Crystal Structure of the Papain-E-64-C Complex Authors: Yamamoto, D. / Matsumoto, K. / Ohishi, H. / Ishida, T. / Inoue, M. / Kitamura, K. / Mizuno, H. | ||||||
| History |
| ||||||
| Remark 700 | SHEET THE SHEET STRUCTURE OF THIS MOLECULE IS BIFURCATED. IN ORDER TO REPRESENT THIS FEATURE IN THE ...SHEET THE SHEET STRUCTURE OF THIS MOLECULE IS BIFURCATED. IN ORDER TO REPRESENT THIS FEATURE IN THE SHEET RECORDS BELOW, TWO SHEETS ARE DEFINED. STRANDS 4, 5, 6 OF S1A ARE IDENTICAL TO STRANDS 2, 3, 4 OF SHEET S1B. |
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1ppp.cif.gz 1ppp.cif.gz | 60.1 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1ppp.ent.gz pdb1ppp.ent.gz | 43.5 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1ppp.json.gz 1ppp.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/pp/1ppp https://data.pdbj.org/pub/pdb/validation_reports/pp/1ppp ftp://data.pdbj.org/pub/pdb/validation_reports/pp/1ppp ftp://data.pdbj.org/pub/pdb/validation_reports/pp/1ppp | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
| ||||||||
| Atom site foot note | 1: CIS PROLINE - PRO 152 2: THE BOND BETWEEN ATOMS O1 AND C2 IS BROKEN WHEN E-64-C BINDS TO PAPAIN IN THIS STRUCTURE. |
- Components
Components
| #1: Protein | Mass: 23449.346 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  |
|---|---|
| #2: Chemical | ChemComp-E6C / |
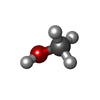
| #3: Chemical | ChemComp-MOH / |
| #4: Water | ChemComp-HOH / |
| Has protein modification | Y |
| Nonpolymer details | E6C IS (2S,3S)-3-(1-(N-(3-METHYLBUTYL)AMINO)- LEUCYLCARBOXYL)OXIRANE-2-CARBOXYLATE. THIS COMPOUND ...E6C IS (2S,3S)-3-(1-(N-(3-METHYLBUTY |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION X-RAY DIFFRACTION |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.36 Å3/Da / Density % sol: 47.94 % | ||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | *PLUS Temperature: 20 ℃ / pH: 9.2 / Method: vapor diffusion, sitting drop | ||||||||||||||||||||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Radiation | Scattering type: x-ray |
|---|---|
| Radiation wavelength | Relative weight: 1 |
| Reflection | *PLUS Highest resolution: 1.9 Å / Num. obs: 14811 / Observed criterion σ(F): 3 / Num. measured all: 21135 / Rmerge(I) obs: 0.087 |
- Processing
Processing
| Software | Name: PROLSQ / Classification: refinement | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Rfactor obs: 0.194 / Highest resolution: 1.9 Å | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Highest resolution: 1.9 Å
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Software | *PLUS Name: PROLSQ / Classification: refinement | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS Highest resolution: 1.9 Å / Lowest resolution: 20 Å / Num. reflection obs: 14811 / Rfactor obs: 0.194 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS |
 Movie
Movie Controller
Controller




















 PDBj
PDBj