[English] 日本語
 Yorodumi
Yorodumi- PDB-1pgv: Structural Genomics of Caenorhabditis elegans: tropomodulin C-ter... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1pgv | ||||||
|---|---|---|---|---|---|---|---|
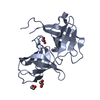
| Title | Structural Genomics of Caenorhabditis elegans: tropomodulin C-terminal domain | ||||||
 Components Components | tropomodulin TMD-1 | ||||||
 Keywords Keywords | PROTEIN BINDING / Structural genomics / tropomodulin / PSI / Protein Structure Initiative / Southeast Collaboratory for Structural Genomics / SECSG | ||||||
| Function / homology |  Function and homology information Function and homology informationSmooth Muscle Contraction / muscle thin filament assembly / pointed-end actin filament capping / nematode larval development / terminal web / myofibril assembly / negative regulation of actin filament depolymerization / embryo development ending in birth or egg hatching / locomotion / sarcomere organization ...Smooth Muscle Contraction / muscle thin filament assembly / pointed-end actin filament capping / nematode larval development / terminal web / myofibril assembly / negative regulation of actin filament depolymerization / embryo development ending in birth or egg hatching / locomotion / sarcomere organization / tropomyosin binding / myofibril / striated muscle thin filament / actin filament organization / actin filament binding / cytoskeleton / cytoplasm Similarity search - Function | ||||||
| Biological species |  | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.8 Å MOLECULAR REPLACEMENT / Resolution: 1.8 Å | ||||||
 Authors Authors | Symersky, J. / Lu, S. / Li, S. / Chen, L. / Meehan, E. / Luo, M. / Qiu, S. / Bunzel, R.J. / Luo, D. / Arabashi, A. ...Symersky, J. / Lu, S. / Li, S. / Chen, L. / Meehan, E. / Luo, M. / Qiu, S. / Bunzel, R.J. / Luo, D. / Arabashi, A. / Nagy, L.A. / Lin, G. / Luan, W.C.-H. / Carson, M. / Gray, R. / Huang, W. / Southeast Collaboratory for Structural Genomics (SECSG) | ||||||
 Citation Citation |  Journal: Proteins / Year: 2004 Journal: Proteins / Year: 2004Title: Structural genomics of Caenorhabditis elegans: crystal structure of the tropomodulin C-terminal domain Authors: Lu, S. / Symersky, J. / Li, S. / Carson, M. / Chen, L. / Meehan, E. / Luo, M. #1:  Journal: Acta Crystallogr.,Sect.D / Year: 2003 Journal: Acta Crystallogr.,Sect.D / Year: 2003Title: Purification, nanocrystallization and preliminary X_ray analysis of a C_terminal part of tropomodulin protein 1, isoform a, from Caenorhabditis elegans Authors: Ding, H. / Qiu, S. / Bunzel, R.J. / Luo, D. / Arabashi, A. / Lu, S. / Symersky, J. / Nagy, L.A. / Delucas, L.J. / Li, S. / Luo, M. #2:  Journal: Biophys.J. / Year: 2002 Journal: Biophys.J. / Year: 2002Title: Crystal structure of the C-terminal half of tropomodulin and structural basis of actin filament pointed-end capping Authors: Krieger, I. / Kostyukova, A. / Yamashita, A. / Nitanai, Y. / Maeda, Y. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1pgv.cif.gz 1pgv.cif.gz | 49.5 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1pgv.ent.gz pdb1pgv.ent.gz | 34.3 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1pgv.json.gz 1pgv.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/pg/1pgv https://data.pdbj.org/pub/pdb/validation_reports/pg/1pgv ftp://data.pdbj.org/pub/pdb/validation_reports/pg/1pgv ftp://data.pdbj.org/pub/pdb/validation_reports/pg/1pgv | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  1io0S S: Starting model for refinement |
|---|---|
| Similar structure data | |
| Other databases |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 22389.314 Da / Num. of mol.: 1 / Fragment: C-terminal domain Source method: isolated from a genetically manipulated source Source: (gene. exp.)   |
|---|---|
| #2: Water | ChemComp-HOH / |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.08 Å3/Da / Density % sol: 40.5 % | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | Temperature: 277 K / Method: vapor diffusion, hanging drop / pH: 8 Details: RESERVOIR: 28% PEG400, 0.1 M KCL, 10 MM MGCL2, 0.1 M TRIS, PH 8. PROTEIN SOLUTION: 13.7 MG/ML IN 10 MM HEPES, PH 7.5. DROPS: 1 MICROLITER RESERVOIR + 1 MICROLITER PROTEIN SOLUTION. VAPOR ...Details: RESERVOIR: 28% PEG400, 0.1 M KCL, 10 MM MGCL2, 0.1 M TRIS, PH 8. PROTEIN SOLUTION: 13.7 MG/ML IN 10 MM HEPES, PH 7.5. DROPS: 1 MICROLITER RESERVOIR + 1 MICROLITER PROTEIN SOLUTION. VAPOR DIFFUSION, HANGING DROP, temperature 277K | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Crystal grow | *PLUS Temperature: 277 K / pH: 7.5 / Method: vapor diffusion, hanging dropDetails: Ding, H., (2003) Acta Crystallogr.,Sect.D, 59, 1106. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  APS APS  / Beamline: 22-ID / Wavelength: 0.984 Å / Beamline: 22-ID / Wavelength: 0.984 Å |
| Detector | Type: MARRESEARCH / Detector: CCD / Date: Feb 16, 2003 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.984 Å / Relative weight: 1 |
| Reflection | Resolution: 1.8→50 Å / Num. obs: 16153 / % possible obs: 96.9 % / Observed criterion σ(I): -1 / Redundancy: 8.8 % / Biso Wilson estimate: 24.3 Å2 / Rsym value: 0.064 / Net I/σ(I): 10.6 |
| Reflection shell | Resolution: 1.8→1.86 Å / Redundancy: 3.2 % / Mean I/σ(I) obs: 3.5 / Num. unique all: 1421 / Rsym value: 0.243 / % possible all: 85.3 |
| Reflection | *PLUS Highest resolution: 1.8 Å / Lowest resolution: 50 Å / Num. obs: 16189 / % possible obs: 96.7 % / Num. measured all: 142014 / Rmerge(I) obs: 0.064 |
| Reflection shell | *PLUS Highest resolution: 1.8 Å / % possible obs: 85.3 % / Rmerge(I) obs: 0.243 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB entry 1IO0 Resolution: 1.8→50 Å / Data cutoff high absF: 10000 / Data cutoff high rms absF: 10000 / Data cutoff low absF: 0 / Isotropic thermal model: isotropic / Cross valid method: THROUGHOUT / σ(F): 0 / Stereochemistry target values: Engh & Huber
| ||||||||||||||||||||||||||||
| Solvent computation | Solvent model: FLAT MODEL / Bsol: 45.83 Å2 / ksol: 0.39 e/Å3 | ||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 25.5 Å2 | ||||||||||||||||||||||||||||
| Refine analyze | Luzzati coordinate error obs: 0.21 Å / Luzzati d res low obs: 5 Å / Luzzati sigma a obs: 0.13 Å | ||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.8→50 Å
| ||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 1.8→1.84 Å / Total num. of bins used: 15 /
| ||||||||||||||||||||||||||||
| Xplor file |
| ||||||||||||||||||||||||||||
| Refinement | *PLUS Highest resolution: 1.8 Å / % reflection Rfree: 5 % | ||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||
| Displacement parameters | *PLUS |
 Movie
Movie Controller
Controller











 PDBj
PDBj
