[English] 日本語
 Yorodumi
Yorodumi- PDB-1omu: SOLUTION STRUCTURE OF OVOMUCOID (THIRD DOMAIN) FROM DOMESTIC TURK... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1omu | ||||||
|---|---|---|---|---|---|---|---|
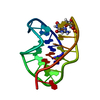
| Title | SOLUTION STRUCTURE OF OVOMUCOID (THIRD DOMAIN) FROM DOMESTIC TURKEY (298K, PH 4.1) (NMR, 50 STRUCTURES) (REFINED MODEL USING NETWORK EDITING ANALYSIS) | ||||||
 Components Components | OVOMUCOID (THIRD DOMAIN) | ||||||
 Keywords Keywords | SERINE PROTEINASE INHIBITOR / SPIN DIFFUSION / NETWORK EDITING / BD-NOESY / CBD-NOESY | ||||||
| Function / homology |  Function and homology information Function and homology informationmolecular function inhibitor activity / serine-type endopeptidase inhibitor activity / extracellular region Similarity search - Function | ||||||
| Biological species |  | ||||||
| Method | SOLUTION NMR | ||||||
 Authors Authors | Hoogstraten, C.G. / Choe, S. / Westler, W.M. / Markley, J.L. | ||||||
 Citation Citation |  Journal: Protein Sci. / Year: 1995 Journal: Protein Sci. / Year: 1995Title: Comparison of the accuracy of protein solution structures derived from conventional and network-edited NOESY data. Authors: Hoogstraten, C.G. / Choe, S. / Westler, W.M. / Markley, J.L. #1:  Journal: J.Mol.Biol. / Year: 1994 Journal: J.Mol.Biol. / Year: 1994Title: Solution Structure of Turkey Ovomucoid Third Domain as Determined from Nuclear Magnetic Resonance Data Authors: Krezel, A.M. / Darba, P. / Robertson, A.D. / Fejzo, J. / Macura, S. / Markley, J.L. #2:  Journal: Protein Eng. / Year: 1993 Journal: Protein Eng. / Year: 1993Title: Overexpression and Purification of Avian Ovomucoid Third Domains in Escherichia Coli Authors: Hinck, A.P. / Walkenhorst, W.F. / Westler, W.M. / Choe, S. / Markley, J.L. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1omu.cif.gz 1omu.cif.gz | 780.2 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1omu.ent.gz pdb1omu.ent.gz | 657.2 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1omu.json.gz 1omu.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/om/1omu https://data.pdbj.org/pub/pdb/validation_reports/om/1omu ftp://data.pdbj.org/pub/pdb/validation_reports/om/1omu ftp://data.pdbj.org/pub/pdb/validation_reports/om/1omu | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| Atom site foot note | 1: CIS PROLINE - PRO 12 | |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein | Mass: 6026.811 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Description: MATERIAL PRODUCED FROM THE NATURAL SOURCE WAS USED FOR A PORTION OF THE DATA. PLASMID PTNOM3 WAS MODIFIED BY MUTATING TWO METHIONINES IN THE SNASE MOIETY TO ALANINES TO FACILITATE PURIFICATION Cell line: BL21 / Cellular location: EGG WHITE / Gene: OM3 / Organ: EGG / Plasmid: PTNOM3 (FUSION WITH STAPH. NUCLEASE) / Gene (production host): OM3 / Production host:  |
|---|---|
| Has protein modification | Y |
-Experimental details
-Experiment
| Experiment | Method: SOLUTION NMR |
|---|
- Sample preparation
Sample preparation
| Crystal grow | *PLUS Method: other |
|---|
- Processing
Processing
| Software |
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR software | Name:  X-PLOR / Version: 3.1 / Developer: BRUNGER / Classification: refinement X-PLOR / Version: 3.1 / Developer: BRUNGER / Classification: refinement | ||||||||||||
| NMR ensemble | Conformers submitted total number: 50 |
 Movie
Movie Controller
Controller





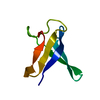




 PDBj
PDBj
