+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1nxb | ||||||
|---|---|---|---|---|---|---|---|
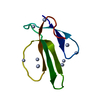
| Title | STRUCTURE AND FUNCTION OF SNAKE VENOM CURARIMIMETIC NEUROTOXINS | ||||||
 Components Components | NEUROTOXIN B | ||||||
 Keywords Keywords | NEUROTOXIN (POST-SYNAPTIC) | ||||||
| Function / homology |  Function and homology information Function and homology informationacetylcholine receptor inhibitor activity / ion channel regulator activity / toxin activity / extracellular region Similarity search - Function | ||||||
| Biological species |  Laticauda semifasciata (broad-banded blue sea krait) Laticauda semifasciata (broad-banded blue sea krait) | ||||||
| Method |  X-RAY DIFFRACTION / Resolution: 1.38 Å X-RAY DIFFRACTION / Resolution: 1.38 Å | ||||||
 Authors Authors | Tsernoglou, D. / Petsko, G.A. | ||||||
 Citation Citation |  Journal: Mol.Pharmacol. / Year: 1978 Journal: Mol.Pharmacol. / Year: 1978Title: Structure and function of snake venom curarimimetic neurotoxins. Authors: Tsernoglou, D. / Petsko, G.A. / Hudson, R.A. #1:  Journal: Science / Year: 1977 Journal: Science / Year: 1977Title: Molecular Graphics. Application to the Structure Determination of a Snake Venom Neurotoxin Authors: Tsernoglou, D. / Petsko, G.A. / Mcqueenjunior, J.E. / Hermans, J. #2:  Journal: Biochim.Biophys.Acta / Year: 1977 Journal: Biochim.Biophys.Acta / Year: 1977Title: Protein Sequencing by Computer Graphics Authors: Tsernoglou, D. / Petsko, G.A. / TU, A.T. #3:  Journal: Proc.Natl.Acad.Sci.USA / Year: 1977 Journal: Proc.Natl.Acad.Sci.USA / Year: 1977Title: Three-Dimensional Structure of Neurotoxin a from Venom of the Philippines Sea Snake Authors: Tsernoglou, D. / Petsko, G.A. #4:  Journal: FEBS Lett. / Year: 1976 Journal: FEBS Lett. / Year: 1976Title: The Crystal Structure of a Post-Synaptic Neurotoxin from Sea Snake at 2.2 Angstroms Resolution Authors: Tsernoglou, D. / Petsko, G.A. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1nxb.cif.gz 1nxb.cif.gz | 32 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1nxb.ent.gz pdb1nxb.ent.gz | 15.3 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1nxb.json.gz 1nxb.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/nx/1nxb https://data.pdbj.org/pub/pdb/validation_reports/nx/1nxb ftp://data.pdbj.org/pub/pdb/validation_reports/nx/1nxb ftp://data.pdbj.org/pub/pdb/validation_reports/nx/1nxb | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 6877.759 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Laticauda semifasciata (broad-banded blue sea krait) Laticauda semifasciata (broad-banded blue sea krait)References: UniProt: Q90VW1 | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| #2: Chemical | | #3: Water | ChemComp-HOH / | Compound details | THE PROTEIN IS PROBABLY IDENTICAL TO ERABUTOXIN FROM JAPANESE SEA SNAKES. THIS POINT IS DISCUSSED ...THE PROTEIN IS PROBABLY IDENTICAL TO ERABUTOXIN | Has protein modification | Y | Nonpolymer details | COORDINATES FOR TWO SULFATE IONS ARE INCLUDED BELOW. THERE IS PROBABLY AT LEAST ONE OTHER BOUND ...COORDINATE | Sequence details | RESIDUE NUMBERING IS SEQUENTIAL. IN PUBLISHED PAPERS A GENERAL HOMOLOGY SEQUENCE NUMBERING IS USED ...RESIDUE NUMBERING IS SEQUENTIAL | |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION X-RAY DIFFRACTION |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 1.8 Å3/Da / Density % sol: 31.64 % |
|---|
-Data collection
| Radiation | Scattering type: x-ray |
|---|---|
| Radiation wavelength | Relative weight: 1 |
- Processing
Processing
| Software | Name: PROLSQ / Classification: refinement | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Highest resolution: 1.38 Å | ||||||||||||
| Refinement step | Cycle: LAST / Highest resolution: 1.38 Å
|
 Movie
Movie Controller
Controller












 PDBj
PDBj




