+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1nvr | ||||||
|---|---|---|---|---|---|---|---|
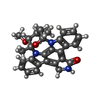
| Title | The Complex Structure Of Checkpoint Kinase Chk1/Staurosporine | ||||||
 Components Components |
| ||||||
 Keywords Keywords | TRANSFERASE / Chk1-Staurosporine complex | ||||||
| Function / homology |  Function and homology information Function and homology informationnegative regulation of mitotic nuclear division / apoptotic process involved in development / negative regulation of G0 to G1 transition / histone H3T11 kinase activity / regulation of mitotic centrosome separation / mitotic G2/M transition checkpoint / peptidyl-threonine phosphorylation / inner cell mass cell proliferation / regulation of double-strand break repair via homologous recombination / nucleus organization ...negative regulation of mitotic nuclear division / apoptotic process involved in development / negative regulation of G0 to G1 transition / histone H3T11 kinase activity / regulation of mitotic centrosome separation / mitotic G2/M transition checkpoint / peptidyl-threonine phosphorylation / inner cell mass cell proliferation / regulation of double-strand break repair via homologous recombination / nucleus organization / negative regulation of gene expression, epigenetic / mitotic G2 DNA damage checkpoint signaling / Transcriptional Regulation by E2F6 / replicative senescence / Presynaptic phase of homologous DNA pairing and strand exchange / Activation of ATR in response to replication stress / Chk1/Chk2(Cds1) mediated inactivation of Cyclin B:Cdk1 complex / signal transduction in response to DNA damage / positive regulation of cell cycle / DNA damage checkpoint signaling / regulation of signal transduction by p53 class mediator / replication fork / condensed nuclear chromosome / TP53 Regulates Transcription of DNA Repair Genes / cellular response to mechanical stimulus / Signaling by SCF-KIT / G2/M DNA damage checkpoint / Ubiquitin-Mediated Degradation of Phosphorylated Cdc25A / G2/M transition of mitotic cell cycle / regulation of cell population proliferation / Processing of DNA double-strand break ends / Regulation of TP53 Activity through Phosphorylation / protein phosphorylation / protein kinase activity / DNA replication / non-specific serine/threonine protein kinase / chromatin remodeling / protein domain specific binding / protein serine kinase activity / DNA repair / protein serine/threonine kinase activity / apoptotic process / DNA damage response / centrosome / chromatin / protein-containing complex / : / nucleoplasm / ATP binding / nucleus / cytoplasm / cytosol Similarity search - Function | ||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.8 Å MOLECULAR REPLACEMENT / Resolution: 1.8 Å | ||||||
 Authors Authors | Zhao, B. / Bower, M.J. / McDevitt, P.J. / Zhao, H. / Davis, S.T. / Johanson, K.O. / Green, S.M. / Concha, N.O. / Zhou, B.B. | ||||||
 Citation Citation |  Journal: J.Biol.Chem. / Year: 2002 Journal: J.Biol.Chem. / Year: 2002Title: Structural Basis for Chk1 Inhibition by UCN-01 Authors: Zhao, B. / Bower, M.J. / McDevitt, P.J. / Zhao, H. / Davis, S.T. / Johanson, K.O. / Green, S.M. / Concha, N.O. / Zhou, B.B. | ||||||
| History |
| ||||||
| Remark 999 | SEQUENCE A five-residue peptide 301-305 was found with the enzyme. Residues were named based on the ...SEQUENCE A five-residue peptide 301-305 was found with the enzyme. Residues were named based on the electron density. The peptide could be the part of C-terminal missing residues. |
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1nvr.cif.gz 1nvr.cif.gz | 73.7 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1nvr.ent.gz pdb1nvr.ent.gz | 53.1 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1nvr.json.gz 1nvr.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/nv/1nvr https://data.pdbj.org/pub/pdb/validation_reports/nv/1nvr ftp://data.pdbj.org/pub/pdb/validation_reports/nv/1nvr ftp://data.pdbj.org/pub/pdb/validation_reports/nv/1nvr | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  1nvqC  1nvsC  2phkS C: citing same article ( S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 33042.988 Da / Num. of mol.: 1 / Fragment: CHK1KD (RESIDUES 1-289) Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Production host: Homo sapiens (human) / Production host:  unidentified baculovirus / Strain (production host): SF9 unidentified baculovirus / Strain (production host): SF9References: UniProt: O14757, Transferases; Transferring phosphorus-containing groups; Phosphotransferases with an alcohol group as acceptor |
|---|---|
| #2: Protein/peptide | Mass: 433.457 Da / Num. of mol.: 1 / Source method: obtained synthetically |
| #3: Chemical | ChemComp-SO4 / |
| #4: Chemical | ChemComp-STU / |
| #5: Water | ChemComp-HOH / |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.54 Å3/Da / Density % sol: 51.23 % | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | Temperature: 293 K / Method: vapor diffusion, sitting drop / pH: 7.5 Details: PEG8000, ammonium sulfate, glycerol, pH 7.5, VAPOR DIFFUSION, SITTING DROP, temperature 293K | ||||||||||||||||||||
| Crystal grow | *PLUS Method: vapor diffusion | ||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  APS APS  / Beamline: 17-ID / Wavelength: 1 Å / Beamline: 17-ID / Wavelength: 1 Å |
| Detector | Type: MARRESEARCH / Detector: CCD / Date: Dec 2, 2000 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1 Å / Relative weight: 1 |
| Reflection | Resolution: 1.8→50 Å / Num. all: 31548 / Num. obs: 31477 / % possible obs: 99.8 % / Observed criterion σ(F): 1 / Observed criterion σ(I): 1 |
| Reflection | *PLUS Highest resolution: 1.8 Å / % possible obs: 100 % / Redundancy: 4.2 % / Num. measured all: 132242 / Rmerge(I) obs: 0.082 |
- Processing
Processing
| Software |
| |||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB ENTRY 2PHK Resolution: 1.8→20 Å / σ(F): 1 / Stereochemistry target values: Engh & Huber
| |||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.8→20 Å
| |||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||
| Refinement | *PLUS Lowest resolution: 20 Å / % reflection Rfree: 6 % / Rfactor Rfree: 0.237 / Rfactor Rwork: 0.213 | |||||||||||||||||||||||||
| Solvent computation | *PLUS | |||||||||||||||||||||||||
| Displacement parameters | *PLUS | |||||||||||||||||||||||||
| Refine LS restraints | *PLUS
|
 Movie
Movie Controller
Controller













 PDBj
PDBj