[English] 日本語
 Yorodumi
Yorodumi- PDB-1neg: Crystal Structure Analysis of N-and C-terminal labeled SH3-domain... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1neg | ||||||
|---|---|---|---|---|---|---|---|
| Title | Crystal Structure Analysis of N-and C-terminal labeled SH3-domain of alpha-Chicken Spectrin | ||||||
 Components Components | Spectrin alpha chain, brain | ||||||
 Keywords Keywords | STRUCTURAL PROTEIN / SH3-domain fold / five antiparallel beta sheets | ||||||
| Function / homology |  Function and homology information Function and homology informationcostamere / actin filament capping / cortical actin cytoskeleton / cell projection / actin filament binding / cell junction / actin cytoskeleton organization / calmodulin binding / calcium ion binding / plasma membrane Similarity search - Function | ||||||
| Biological species |  | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  MOLECULAR REPLACEMENT / Resolution: 2.3 Å MOLECULAR REPLACEMENT / Resolution: 2.3 Å | ||||||
 Authors Authors | Mueller, U. / Buessow, K. / Diehl, A. / Niesen, F.H. / Nyarsik, L. / Heinemann, U. | ||||||
 Citation Citation |  Journal: J.STRUCT.FUNCT.GENOM. / Year: 2003 Journal: J.STRUCT.FUNCT.GENOM. / Year: 2003Title: Rapid purification and crystal structure analysis of a small protein carrying two terminal affinity tags Authors: Mueller, U. / Buessow, K. / Diehl, A. / Bartl, F.J. / Niesen, F.H. / Nyarsik, L. / Heinemann, U. | ||||||
| History |
| ||||||
| Remark 999 | SEQUENCE ACCORDING TO THE AUTHOR, THIS IS PART OF A STREP2 TAG. |
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1neg.cif.gz 1neg.cif.gz | 27.5 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1neg.ent.gz pdb1neg.ent.gz | 17.2 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1neg.json.gz 1neg.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/ne/1neg https://data.pdbj.org/pub/pdb/validation_reports/ne/1neg ftp://data.pdbj.org/pub/pdb/validation_reports/ne/1neg ftp://data.pdbj.org/pub/pdb/validation_reports/ne/1neg | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  1shgS S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 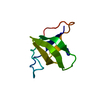
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
| ||||||||
| Components on special symmetry positions |
|
- Components
Components
| #1: Protein | Mass: 9643.871 Da / Num. of mol.: 1 / Fragment: SH3-domain Source method: isolated from a genetically manipulated source Source: (gene. exp.)   |
|---|---|
| #2: Chemical | ChemComp-AZI / |
| #3: Water | ChemComp-HOH / |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.44 Å3/Da / Density % sol: 49.16 % |
|---|---|
| Crystal grow | Temperature: 293 K / Method: vapor diffusion, hanging drop / pH: 6.5 Details: Dioxane, MES, Ammonium sulfate, pH 6.5, VAPOR DIFFUSION, HANGING DROP, temperature 293K |
| Crystal grow | *PLUS Method: vapor diffusion, hanging drop |
-Data collection
| Diffraction | Mean temperature: 110 K |
|---|---|
| Diffraction source | Source:  ROTATING ANODE / Type: RIGAKU RU200 / Wavelength: 1.5418 Å ROTATING ANODE / Type: RIGAKU RU200 / Wavelength: 1.5418 Å |
| Detector | Type: MARRESEARCH / Detector: IMAGE PLATE / Date: Nov 7, 2000 / Details: mirrors |
| Radiation | Monochromator: Supper mirrors / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.5418 Å / Relative weight: 1 |
| Reflection | Resolution: 2.3→31.1 Å / Num. all: 4077 / Num. obs: 3858 / % possible obs: 99.2 % / Observed criterion σ(F): 0 / Observed criterion σ(I): 0 / Redundancy: 4.27 % / Biso Wilson estimate: 26.6 Å2 / Rsym value: 0.109 / Net I/σ(I): 13.05 |
| Reflection shell | Resolution: 2.3→2.33 Å / Mean I/σ(I) obs: 1.94 / Num. unique all: 130 / Rsym value: 0.461 / % possible all: 95.6 |
| Reflection | *PLUS Lowest resolution: 31 Å / Redundancy: 4.4 % / Rmerge(I) obs: 0.109 |
| Reflection shell | *PLUS % possible obs: 95.6 % / Rmerge(I) obs: 0.461 / Mean I/σ(I) obs: 1.92 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 1SHG Resolution: 2.3→31.09 Å / Rfactor Rfree error: 0.013 / Isotropic thermal model: RESTRAINED / Cross valid method: THROUGHOUT / σ(F): 0 / σ(I): 0 / Stereochemistry target values: Engh & Huber / Details: BULK SOLVENT MODEL USED
| ||||||||||||||||||||||||||||||||||||
| Solvent computation | Solvent model: FLAT MODEL / Bsol: 48.9616 Å2 / ksol: 0.410835 e/Å3 | ||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 24.8 Å2
| ||||||||||||||||||||||||||||||||||||
| Refine analyze | Luzzati coordinate error free: 0.35 Å / Luzzati sigma a free: 0.31 Å | ||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.3→31.09 Å
| ||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 2.3→2.44 Å / Rfactor Rfree error: 0.041 / Total num. of bins used: 6
| ||||||||||||||||||||||||||||||||||||
| Xplor file |
| ||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS Highest resolution: 2.3 Å / Lowest resolution: 31.1 Å / % reflection Rfree: 10 % / Rfactor obs: 0.2 / Rfactor Rwork: 0.2 | ||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS | ||||||||||||||||||||||||||||||||||||
| Refine LS restraints | *PLUS
|
 Movie
Movie Controller
Controller












 PDBj
PDBj


