[English] 日本語
 Yorodumi
Yorodumi- PDB-1mzv: Crystal Structure of Adenine Phosphoribosyltransferase (APRT) Fro... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1mzv | ||||||
|---|---|---|---|---|---|---|---|
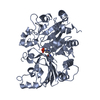
| Title | Crystal Structure of Adenine Phosphoribosyltransferase (APRT) From Leishmania tarentolae | ||||||
 Components Components | Adenine Phosphoribosyltransferase | ||||||
 Keywords Keywords | TRANSFERASE / alpha/beta structure | ||||||
| Function / homology |  Function and homology information Function and homology informationadenine phosphoribosyltransferase / adenine phosphoribosyltransferase activity / purine ribonucleoside salvage / nucleotide binding / cytoplasm Similarity search - Function | ||||||
| Biological species |  Leishmania tarentolae (eukaryote) Leishmania tarentolae (eukaryote) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2.2 Å MOLECULAR REPLACEMENT / Resolution: 2.2 Å | ||||||
 Authors Authors | Thiemann, O.H. / Silva, M. / Oliva, G. / Silva, C.H.T.P. / Iulek, J. | ||||||
 Citation Citation |  Journal: Biochim.Biophys.Acta / Year: 2004 Journal: Biochim.Biophys.Acta / Year: 2004Title: Crystal structure of adenine phosphoribosyltransferase from Leishmania tarentolae: potential implications for APRT catalytic mechanism. Authors: Silva, M. / Silva, C.H.T.P. / Iulek, J. / Oliva, G. / Thiemann, O.H. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1mzv.cif.gz 1mzv.cif.gz | 61.8 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1mzv.ent.gz pdb1mzv.ent.gz | 44.9 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1mzv.json.gz 1mzv.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/mz/1mzv https://data.pdbj.org/pub/pdb/validation_reports/mz/1mzv ftp://data.pdbj.org/pub/pdb/validation_reports/mz/1mzv ftp://data.pdbj.org/pub/pdb/validation_reports/mz/1mzv | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  1qb7S S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| Unit cell |
| ||||||||
| Details | The biological assembly is a dimer generated from the monomer in the asymmetric unit by the operation: y-x, y, 1/2-z |
- Components
Components
| #1: Protein | Mass: 25913.826 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Leishmania tarentolae (eukaryote) / Plasmid: pET29(+) / Production host: Leishmania tarentolae (eukaryote) / Plasmid: pET29(+) / Production host:  References: UniProt: O77103, adenine phosphoribosyltransferase |
|---|---|
| #2: Chemical | ChemComp-PO4 / |
| #3: Chemical | ChemComp-AMP / |
| #4: Water | ChemComp-HOH / |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.72 Å3/Da / Density % sol: 54.75 % | ||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | Temperature: 277 K / pH: 7 Details: MOPS 100 mM, 22% isopropanol, Protein at 8 mg/ml, pH 7.00, temperature 277K | ||||||||||||||||||||||||||||||
| Crystal grow | *PLUS Temperature: 4 ℃ / pH: 7 / Method: vapor diffusion, hanging drop | ||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  LNLS LNLS  / Beamline: D03B-MX1 / Wavelength: 1.38 / Beamline: D03B-MX1 / Wavelength: 1.38 |
| Detector | Type: MARRESEARCH / Detector: IMAGE PLATE / Date: Aug 14, 1999 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.38 Å / Relative weight: 1 |
| Reflection | Resolution: 2.2→28.27 Å / Num. obs: 15513 / % possible obs: 99.5 % / Observed criterion σ(I): 0 |
| Reflection shell | Resolution: 2.2→2.24 Å / % possible all: 93.1 |
| Reflection | *PLUS Highest resolution: 2.2 Å / Lowest resolution: 30 Å / Num. obs: 14724 / % possible obs: 99.5 % / Num. measured all: 229639 / Rmerge(I) obs: 0.06 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB ENTRY 1QB7 Resolution: 2.2→28.27 Å / Cor.coef. Fo:Fc: 0.958 / Cor.coef. Fo:Fc free: 0.945 / TLS residual ADP flag: LIKELY RESIDUAL / Cross valid method: THROUGHOUT / σ(F): 0 / Details: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Ion probe radii: 0.8 Å / Shrinkage radii: 0.8 Å / VDW probe radii: 1.4 Å / Solvent model: BABINET MODEL WITH MASK | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 24.16 Å2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.2→28.27 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 2.2→2.26 Å / Total num. of bins used: 20 /
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS params. | Method: refined / Origin x: 8.3269 Å / Origin y: 31.2102 Å / Origin z: 68.421 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS group |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS Rfactor Rfree: 0.2178 / Rfactor Rwork: 0.1809 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints | *PLUS
|
 Movie
Movie Controller
Controller











 PDBj
PDBj










