[English] 日本語
 Yorodumi
Yorodumi- PDB-1m77: Near Atomic Resolution Crystal Structure of an A-DNA Decamer d(CC... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1m77 | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Near Atomic Resolution Crystal Structure of an A-DNA Decamer d(CCCGATCGGG): Cobalt Hexammine Interactions with A-DNA | ||||||||||||||||||
 Components Components | 5'-D(* Keywords KeywordsDNA / A-DNA / Cobalt hexammine | Function / homology | COBALT HEXAMMINE(III) / DNA |  Function and homology information Function and homology informationMethod |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  MOLECULAR REPLACEMENT / Resolution: 1.25 Å MOLECULAR REPLACEMENT / Resolution: 1.25 Å  Authors AuthorsRamakrishnan, B. / Sekharudu, C. / Pan, B.C. / Sundaralingam, M. |  Citation Citation Journal: Acta Crystallogr.,Sect.D / Year: 2003 Journal: Acta Crystallogr.,Sect.D / Year: 2003Title: Near-atomic resolution crystal structure of an A-DNA decamer d(CCCGATCGGG): cobalt hexammine interaction with A-DNA. Authors: Ramakrishnan, B. / Sekharudu, C. / Pan, B. / Sundaralingam, M. History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1m77.cif.gz 1m77.cif.gz | 15.5 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1m77.ent.gz pdb1m77.ent.gz | 8.9 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1m77.json.gz 1m77.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/m7/1m77 https://data.pdbj.org/pub/pdb/validation_reports/m7/1m77 ftp://data.pdbj.org/pub/pdb/validation_reports/m7/1m77 ftp://data.pdbj.org/pub/pdb/validation_reports/m7/1m77 | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 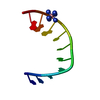
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| Unit cell |
|
- Components
Components
| #1: DNA chain | Mass: 3045.992 Da / Num. of mol.: 1 / Source method: obtained synthetically |
|---|---|
| #2: Chemical | ChemComp-NCO / |
| #3: Water | ChemComp-HOH / |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 1.91 Å3/Da / Density % sol: 35.5 % | ||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | Temperature: 298 K / Method: vapor diffusion, hanging drop / pH: 6 Details: cobalt hexammine, cacodylate, pH 6.0, VAPOR DIFFUSION, HANGING DROP, temperature 298K | ||||||||||||||||||||||||||||||
| Components of the solutions |
| ||||||||||||||||||||||||||||||
| Crystal grow | *PLUS | ||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 298 K |
|---|---|
| Diffraction source | Source:  ROTATING ANODE / Type: SIEMENS / Wavelength: 1.5418 Å ROTATING ANODE / Type: SIEMENS / Wavelength: 1.5418 Å |
| Detector | Type: NICOLET / Detector: AREA DETECTOR / Date: Sep 22, 1993 / Details: mirrors |
| Radiation | Monochromator: graphite / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.5418 Å / Relative weight: 1 |
| Reflection | Resolution: 1.25→28 Å / Num. all: 7785 / Num. obs: 6929 / % possible obs: 90 % / Observed criterion σ(F): 1 / Observed criterion σ(I): 1 / Redundancy: 7.4 % / Rsym value: 0.055 |
| Reflection shell | Resolution: 1.25→1.31 Å / Num. unique all: 433 |
| Reflection | *PLUS % possible obs: 90 % / Num. measured all: 51067 / Rmerge(I) obs: 0.055 |
- Processing
Processing
| Software |
| ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: fiber A-DNA Resolution: 1.25→8 Å / Isotropic thermal model: isotropic / σ(F): 2
| ||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.25→8 Å
| ||||||||||||||||||||
| LS refinement shell | Resolution: 1.25→1.31 Å /
| ||||||||||||||||||||
| Refine LS restraints | *PLUS
|
 Movie
Movie Controller
Controller






 PDBj
PDBj