Entry Database : PDB / ID : 4rhdTitle DNA Duplex with Novel ZP Base Pair DNA 9mer novel P nucleobase DNA 9mer novel Z nucleobase Keywords / Function / homology Biological species synthetic construct (others) Method / / / Resolution : 1.7 Å Authors Zhang, W. / Zhang, L. / Benner, S. / Huang, Z. Journal : J.Am.Chem.Soc. / Year : 2015Title : Evolution of functional six-nucleotide DNA.Authors: Zhang, L. / Yang, Z. / Sefah, K. / Bradley, K.M. / Hoshika, S. / Kim, M.J. / Kim, H.J. / Zhu, G. / Jimenez, E. / Cansiz, S. / Teng, I.T. / Champanhac, C. / McLendon, C. / Liu, C. / Zhang, W. ... Authors : Zhang, L. / Yang, Z. / Sefah, K. / Bradley, K.M. / Hoshika, S. / Kim, M.J. / Kim, H.J. / Zhu, G. / Jimenez, E. / Cansiz, S. / Teng, I.T. / Champanhac, C. / McLendon, C. / Liu, C. / Zhang, W. / Gerloff, D.L. / Huang, Z. / Tan, W. / Benner, S.A. History Deposition Oct 1, 2014 Deposition site / Processing site Revision 1.0 Jul 8, 2015 Provider / Type Revision 1.1 Jul 15, 2015 Group Revision 1.2 May 18, 2016 Group Revision 1.3 Feb 28, 2024 Group / Database references / Derived calculationsCategory chem_comp_atom / chem_comp_bond ... chem_comp_atom / chem_comp_bond / database_2 / pdbx_struct_conn_angle / struct_conn / struct_site Item _database_2.pdbx_DOI / _database_2.pdbx_database_accession ... _database_2.pdbx_DOI / _database_2.pdbx_database_accession / _pdbx_struct_conn_angle.ptnr1_auth_asym_id / _pdbx_struct_conn_angle.ptnr1_auth_comp_id / _pdbx_struct_conn_angle.ptnr1_auth_seq_id / _pdbx_struct_conn_angle.ptnr1_label_asym_id / _pdbx_struct_conn_angle.ptnr1_label_atom_id / _pdbx_struct_conn_angle.ptnr1_label_comp_id / _pdbx_struct_conn_angle.ptnr1_label_seq_id / _pdbx_struct_conn_angle.ptnr3_auth_asym_id / _pdbx_struct_conn_angle.ptnr3_auth_comp_id / _pdbx_struct_conn_angle.ptnr3_auth_seq_id / _pdbx_struct_conn_angle.ptnr3_label_asym_id / _pdbx_struct_conn_angle.ptnr3_label_atom_id / _pdbx_struct_conn_angle.ptnr3_label_comp_id / _pdbx_struct_conn_angle.ptnr3_label_seq_id / _struct_conn.pdbx_dist_value / _struct_conn.pdbx_leaving_atom_flag / _struct_conn.ptnr1_auth_asym_id / _struct_conn.ptnr1_auth_comp_id / _struct_conn.ptnr1_auth_seq_id / _struct_conn.ptnr1_label_asym_id / _struct_conn.ptnr1_label_atom_id / _struct_conn.ptnr1_label_comp_id / _struct_conn.ptnr1_label_seq_id / _struct_conn.ptnr2_auth_asym_id / _struct_conn.ptnr2_label_asym_id / _struct_site.pdbx_auth_asym_id / _struct_site.pdbx_auth_comp_id / _struct_site.pdbx_auth_seq_id
Show all Show less
 Open data
Open data Basic information
Basic information Components
Components Keywords
Keywords Function and homology information
Function and homology information X-RAY DIFFRACTION /
X-RAY DIFFRACTION /  SYNCHROTRON /
SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.7 Å
MOLECULAR REPLACEMENT / Resolution: 1.7 Å  Authors
Authors Citation
Citation Journal: J.Am.Chem.Soc. / Year: 2015
Journal: J.Am.Chem.Soc. / Year: 2015 Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 4rhd.cif.gz
4rhd.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb4rhd.ent.gz
pdb4rhd.ent.gz PDB format
PDB format 4rhd.json.gz
4rhd.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads https://data.pdbj.org/pub/pdb/validation_reports/rh/4rhd
https://data.pdbj.org/pub/pdb/validation_reports/rh/4rhd ftp://data.pdbj.org/pub/pdb/validation_reports/rh/4rhd
ftp://data.pdbj.org/pub/pdb/validation_reports/rh/4rhd Links
Links Assembly
Assembly

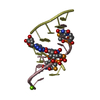

 Components
Components X-RAY DIFFRACTION / Number of used crystals: 1
X-RAY DIFFRACTION / Number of used crystals: 1  Sample preparation
Sample preparation SYNCHROTRON / Site:
SYNCHROTRON / Site:  SSRL
SSRL  / Beamline: BL9-1 / Wavelength: 0.98 Å
/ Beamline: BL9-1 / Wavelength: 0.98 Å Processing
Processing MOLECULAR REPLACEMENT / Resolution: 1.7→87.05 Å / Cor.coef. Fo:Fc: 0.923 / Cor.coef. Fo:Fc free: 0.935 / SU B: 0.007 / SU ML: 0 / Cross valid method: THROUGHOUT / ESU R: 0.08 / ESU R Free: 0.081 / Stereochemistry target values: MAXIMUM LIKELIHOOD / Details: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS
MOLECULAR REPLACEMENT / Resolution: 1.7→87.05 Å / Cor.coef. Fo:Fc: 0.923 / Cor.coef. Fo:Fc free: 0.935 / SU B: 0.007 / SU ML: 0 / Cross valid method: THROUGHOUT / ESU R: 0.08 / ESU R Free: 0.081 / Stereochemistry target values: MAXIMUM LIKELIHOOD / Details: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS Movie
Movie Controller
Controller







 PDBj
PDBj



