+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1iqb | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
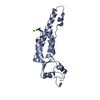
| Title | Crystal Structure of Urtica dioica Agglutinin Isolectin I | |||||||||
 Components Components | AGGLUTININ ISOLECTIN I | |||||||||
 Keywords Keywords | SUGAR BINDING PROTEIN / TWO HOMOLOGOUS HEVEIN-LIKE DOMAINS / ZINC COMPLEX / HOMO-DIMER | |||||||||
| Function / homology |  Function and homology information Function and homology informationendochitinase activity / chitinase / chitin catabolic process / chitin binding / polysaccharide catabolic process / defense response to fungus / cell wall macromolecule catabolic process / carbohydrate binding / killing of cells of another organism / metal ion binding Similarity search - Function | |||||||||
| Biological species |  Urtica dioica (great nettle) Urtica dioica (great nettle) | |||||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  MOLECULAR REPLACEMENT / Resolution: 1.9 Å MOLECULAR REPLACEMENT / Resolution: 1.9 Å | |||||||||
 Authors Authors | Harata, K. / Schubert, W.D. / Muraki, M. | |||||||||
 Citation Citation |  Journal: Acta Crystallogr.,Sect.D / Year: 2001 Journal: Acta Crystallogr.,Sect.D / Year: 2001Title: Structure of Urtica dioica agglutinin isolectin I: dimer formation mediated by two zinc ions bound at the sugar-binding site. Authors: Harata, K. / Schubert, W.D. / Muraki, M. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1iqb.cif.gz 1iqb.cif.gz | 47.1 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1iqb.ent.gz pdb1iqb.ent.gz | 33.2 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1iqb.json.gz 1iqb.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/iq/1iqb https://data.pdbj.org/pub/pdb/validation_reports/iq/1iqb ftp://data.pdbj.org/pub/pdb/validation_reports/iq/1iqb ftp://data.pdbj.org/pub/pdb/validation_reports/iq/1iqb | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  1ehdS S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 9408.387 Da / Num. of mol.: 2 / Fragment: residues 1-89 / Mutation: Q1(PCA) / Source method: isolated from a natural source / Source: (natural)  Urtica dioica (great nettle) / References: UniProt: P11218 Urtica dioica (great nettle) / References: UniProt: P11218#2: Chemical | #3: Water | ChemComp-HOH / | Has protein modification | Y | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.11 Å3/Da / Density % sol: 41.6 % | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | Temperature: 298 K / Method: vapor diffusion, hanging drop / pH: 6.5 Details: PEG8000, zinc acetate, cacodylate, pH 6.5, VAPOR DIFFUSION, HANGING DROP, temperature 298K | ||||||||||||||||||||
| Crystal grow | *PLUS | ||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 286 K |
|---|---|
| Diffraction source | Source:  ROTATING ANODE / Type: ENRAF-NONIUS FR571 / Wavelength: 1.5418 Å ROTATING ANODE / Type: ENRAF-NONIUS FR571 / Wavelength: 1.5418 Å |
| Detector | Type: ENRAF-NONIUS FAST / Detector: AREA DETECTOR / Date: Feb 1, 1998 |
| Radiation | Monochromator: graphite / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.5418 Å / Relative weight: 1 |
| Reflection | Resolution: 1.73→25.1 Å / Num. all: 16600 / Num. obs: 14774 / % possible obs: 89 % / Observed criterion σ(F): 0 / Observed criterion σ(I): 0 / Redundancy: 3.2 % / Rmerge(I) obs: 0.069 |
| Reflection shell | Resolution: 1.73→1.76 Å / Rmerge(I) obs: 0.294 / % possible all: 25 |
| Reflection | *PLUS Num. measured all: 47712 |
- Processing
Processing
| Software |
| |||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB ENTRY 1EHD Resolution: 1.9→8 Å / Isotropic thermal model: Isotropic / σ(F): 2 / Stereochemistry target values: Engh & Huber
| |||||||||||||||||||||||||
| Displacement parameters | Biso mean: 27.5 Å2 | |||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.9→8 Å
| |||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||
| LS refinement shell | Resolution: 1.9→1.93 Å
| |||||||||||||||||||||||||
| Software | *PLUS Name:  X-PLOR / Version: 3.1 / Classification: refinement X-PLOR / Version: 3.1 / Classification: refinement | |||||||||||||||||||||||||
| Refinement | *PLUS Lowest resolution: 8 Å / σ(F): 2 / % reflection Rfree: 13 % / Rfactor obs: 0.205 / Rfactor Rfree: 0.26 | |||||||||||||||||||||||||
| Solvent computation | *PLUS | |||||||||||||||||||||||||
| Displacement parameters | *PLUS Biso mean: 27.5 Å2 | |||||||||||||||||||||||||
| LS refinement shell | *PLUS Rfactor Rfree: 0.387 / Rfactor Rwork: 0.305 |
 Movie
Movie Controller
Controller












 PDBj
PDBj


