[English] 日本語
 Yorodumi
Yorodumi- PDB-1iq5: Calmodulin/nematode CA2+/Calmodulin dependent kinase kinase fragment -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1iq5 | ||||||
|---|---|---|---|---|---|---|---|
| Title | Calmodulin/nematode CA2+/Calmodulin dependent kinase kinase fragment | ||||||
 Components Components |
| ||||||
 Keywords Keywords | METAL BINDING PROTEIN/PROTEIN BINDING / EF-hand / CALMODULIN / PROTEIN-PEPTIDE COMPLEX / METAL BINDING PROTEIN-PROTEIN BINDING COMPLEX | ||||||
| Function / homology |  Function and homology information Function and homology informationCaMK IV-mediated phosphorylation of CREB / CREB1 phosphorylation through the activation of CaMKII/CaMKK/CaMKIV cascasde / Activation of RAC1 downstream of NMDARs / Ca2+/calmodulin-dependent protein kinase / CAMKK-AMPK signaling cascade / calcium/calmodulin-dependent protein kinase activity / myosin II complex / calcium-mediated signaling / calmodulin binding / signaling receptor binding ...CaMK IV-mediated phosphorylation of CREB / CREB1 phosphorylation through the activation of CaMKII/CaMKK/CaMKIV cascasde / Activation of RAC1 downstream of NMDARs / Ca2+/calmodulin-dependent protein kinase / CAMKK-AMPK signaling cascade / calcium/calmodulin-dependent protein kinase activity / myosin II complex / calcium-mediated signaling / calmodulin binding / signaling receptor binding / protein serine kinase activity / protein serine/threonine kinase activity / calcium ion binding / ATP binding / metal ion binding / nucleus / cytoplasm Similarity search - Function | ||||||
| Biological species | |||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.8 Å MOLECULAR REPLACEMENT / Resolution: 1.8 Å | ||||||
 Authors Authors | Kurokawa, H. / Osawa, M. / Kurihara, H. / Katayama, N. / Tokumitsu, H. / Swindells, M.B. / Kainosho, M. / Ikura, M. | ||||||
 Citation Citation |  Journal: J.Mol.Biol. / Year: 2001 Journal: J.Mol.Biol. / Year: 2001Title: Target-induced conformational adaptation of calmodulin revealed by the crystal structure of a complex with nematode Ca(2+)/calmodulin-dependent kinase kinase peptide Authors: Kurokawa, H. / Osawa, M. / Kurihara, H. / Katayama, N. / Tokumitsu, H. / Swindells, M.B. / Kainosho, M. / Ikura, M. #1:  Journal: Nat.Struct.Biol. / Year: 1999 Journal: Nat.Struct.Biol. / Year: 1999Title: A novel target recognition revealed by calmodulin in complex with Ca2+-calmodulin-dependent kinase kinase Authors: Osawa, M. / Tokumitsu, H. / Swindells, M.B. / Kurihara, H. / Orita, M. / Shibanuma, T. / Furuya, T. / Ikura, M. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1iq5.cif.gz 1iq5.cif.gz | 47.9 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1iq5.ent.gz pdb1iq5.ent.gz | 33.7 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1iq5.json.gz 1iq5.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/iq/1iq5 https://data.pdbj.org/pub/pdb/validation_reports/iq/1iq5 ftp://data.pdbj.org/pub/pdb/validation_reports/iq/1iq5 ftp://data.pdbj.org/pub/pdb/validation_reports/iq/1iq5 | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 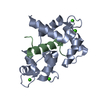
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 16852.545 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  | ||
|---|---|---|---|
| #2: Protein/peptide | Mass: 3198.920 Da / Num. of mol.: 1 / Fragment: CALMODULIN BINDING DOMAIN (RESIDUES 331-357) / Source method: obtained synthetically Details: This sequence occurs naturally in Caenorhabditis elegans References: UniProt: Q8T8D4, UniProt: Q3Y416*PLUS, Transferases; Transferring phosphorus-containing groups; Phosphotransferases with an alcohol group as acceptor | ||
| #3: Chemical | ChemComp-CA / #4: Water | ChemComp-HOH / | |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.4 Å3/Da / Density % sol: 48.81 % | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | Temperature: 293 K / Method: vapor diffusion, hanging drop / pH: 5 Details: ammonium sulfate, citratet, sodium tartrae, pH 5.0, VAPOR DIFFUSION, HANGING DROP, temperature 293K | ||||||||||||||||||||
| Crystal grow | *PLUS Method: vapor diffusion | ||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 293 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  Photon Factory Photon Factory  / Beamline: BL-6B / Wavelength: 1 Å / Beamline: BL-6B / Wavelength: 1 Å |
| Detector | Type: CUSTOM-MADE / Detector: IMAGE PLATE / Date: Feb 10, 1999 |
| Radiation | Monochromator: Si 111 CHANNEL / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1 Å / Relative weight: 1 |
| Reflection | Resolution: 1.8→100 Å / Num. all: 57477 / Num. obs: 17155 / % possible obs: 94.6 % / Observed criterion σ(I): 2 / Redundancy: 3 % / Biso Wilson estimate: 25.798 Å2 / Rmerge(I) obs: 0.024 / Net I/σ(I): 21.9 |
| Reflection shell | Resolution: 1.8→1.91 Å / Redundancy: 2.6 % / Rmerge(I) obs: 0.212 / % possible all: 89.1 |
| Reflection | *PLUS Lowest resolution: 100 Å / Num. measured all: 57477 |
| Reflection shell | *PLUS % possible obs: 89.1 % |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT / Resolution: 1.8→100 Å / σ(I): 2 / Stereochemistry target values: Engh & Huber MOLECULAR REPLACEMENT / Resolution: 1.8→100 Å / σ(I): 2 / Stereochemistry target values: Engh & Huber
| ||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.8→100 Å
| ||||||||||||||||||||||||||||||||||||
| Software | *PLUS Name:  X-PLOR / Version: 3.851 / Classification: refinement X-PLOR / Version: 3.851 / Classification: refinement | ||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS Highest resolution: 1.8 Å / Lowest resolution: 100 Å / Rfactor obs: 0.217 | ||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS | ||||||||||||||||||||||||||||||||||||
| Refine LS restraints | *PLUS
|
 Movie
Movie Controller
Controller












 PDBj
PDBj



