[English] 日本語
 Yorodumi
Yorodumi- PDB-1ilq: CXCR-1 N-TERMINAL PEPTIDE BOUND TO INTERLEUKIN-8 (MINIMIZED MEAN) -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1ilq | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
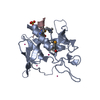
| Title | CXCR-1 N-TERMINAL PEPTIDE BOUND TO INTERLEUKIN-8 (MINIMIZED MEAN) | |||||||||
 Components Components |
| |||||||||
 Keywords Keywords | CYTOKINE | |||||||||
| Function / homology |  Function and homology information Function and homology informationregulation of single stranded viral RNA replication via double stranded DNA intermediate / regulation of entry of bacterium into host cell / interleukin-8 receptor binding / interleukin-8 receptor activity / interleukin-8 binding / negative regulation of cell adhesion molecule production / chemokine receptor activity / ATF4 activates genes in response to endoplasmic reticulum stress / CXCR chemokine receptor binding / embryonic digestive tract development ...regulation of single stranded viral RNA replication via double stranded DNA intermediate / regulation of entry of bacterium into host cell / interleukin-8 receptor binding / interleukin-8 receptor activity / interleukin-8 binding / negative regulation of cell adhesion molecule production / chemokine receptor activity / ATF4 activates genes in response to endoplasmic reticulum stress / CXCR chemokine receptor binding / embryonic digestive tract development / neutrophil activation / induction of positive chemotaxis / C-C chemokine receptor activity / C-C chemokine binding / positive regulation of neutrophil chemotaxis / chemokine activity / Chemokine receptors bind chemokines / negative regulation of G protein-coupled receptor signaling pathway / dendritic cell chemotaxis / Interleukin-10 signaling / cellular response to interleukin-1 / regulation of cell adhesion / neutrophil chemotaxis / cellular response to fibroblast growth factor stimulus / secretory granule membrane / response to endoplasmic reticulum stress / Peptide ligand-binding receptors / calcium-mediated signaling / response to molecule of bacterial origin / receptor internalization / G protein-coupled receptor activity / cellular response to tumor necrosis factor / chemotaxis / positive regulation of angiogenesis / antimicrobial humoral immune response mediated by antimicrobial peptide / heparin binding / positive regulation of cytosolic calcium ion concentration / cellular response to lipopolysaccharide / Senescence-Associated Secretory Phenotype (SASP) / Interleukin-4 and Interleukin-13 signaling / angiogenesis / G alpha (i) signalling events / cell surface receptor signaling pathway / intracellular signal transduction / immune response / G protein-coupled receptor signaling pathway / inflammatory response / external side of plasma membrane / negative regulation of cell population proliferation / negative regulation of gene expression / Neutrophil degranulation / positive regulation of gene expression / signal transduction / : / extracellular region / membrane / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | SOLUTION NMR / RESTRAINED MOLECULAR DYNAMICS | |||||||||
 Authors Authors | Skelton, N.J. / Quan, C. / Lowman, H. | |||||||||
 Citation Citation |  Journal: Structure Fold.Des. / Year: 1999 Journal: Structure Fold.Des. / Year: 1999Title: Structure of a CXC chemokine-receptor fragment in complex with interleukin-8. Authors: Skelton, N.J. / Quan, C. / Reilly, D. / Lowman, H. #1:  Journal: Bioorg.Med.Chem.Lett. / Year: 1997 Journal: Bioorg.Med.Chem.Lett. / Year: 1997Title: Peptide Based Inhibitors of Il-8: Structural Simplification and Improved Potency Authors: Attwood, M.R. / al., et #2:  Journal: Biochemistry / Year: 1990 Journal: Biochemistry / Year: 1990Title: Three-Dimensional Structure of Il-8 in Solution Authors: Clore, G.M. / Appella, E. / Yamada, M. / Gronenborn, A.M. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1ilq.cif.gz 1ilq.cif.gz | 69.9 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1ilq.ent.gz pdb1ilq.ent.gz | 52.8 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1ilq.json.gz 1ilq.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/il/1ilq https://data.pdbj.org/pub/pdb/validation_reports/il/1ilq ftp://data.pdbj.org/pub/pdb/validation_reports/il/1ilq ftp://data.pdbj.org/pub/pdb/validation_reports/il/1ilq | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein | Mass: 8401.807 Da / Num. of mol.: 2 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Plasmid: PPS0170 / Species (production host): Escherichia coli / Cellular location (production host): PERIPLASM / Production host: Homo sapiens (human) / Plasmid: PPS0170 / Species (production host): Escherichia coli / Cellular location (production host): PERIPLASM / Production host:  #2: Protein/peptide | | Mass: 2068.220 Da / Num. of mol.: 1 / Fragment: 9-29 / Mutation: YES Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / References: UniProt: P25024 Homo sapiens (human) / References: UniProt: P25024Has protein modification | Y | |
|---|
-Experimental details
-Experiment
| Experiment | Method: SOLUTION NMR | ||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
| ||||||||||||||||||||||||||||
| NMR details | Text: THE ASSIGNMENTS WERE MADE USING TRIPLE RESONANCE NMR EXPERIMENTS CONDUCTED ON 13C/15N LABELED IL-8 BOUND TO UNLABELED CXCR-1 PEPTIDE (SEE JRNL ENTRY FOR MORE DETAILS) NOE RESTRAINTS WERE ...Text: THE ASSIGNMENTS WERE MADE USING TRIPLE RESONANCE NMR EXPERIMENTS CONDUCTED ON 13C/15N LABELED IL-8 BOUND TO UNLABELED CXCR-1 PEPTIDE (SEE JRNL ENTRY FOR MORE DETAILS) NOE RESTRAINTS WERE OBTAINED FROM 15N EDITED EXPERIMENTS (INTRA IL8), 13C OR 15N FILTERED EXPERIMENTS (INTRA CXCR-1) OR 13C-FILTERED/ EDITED EXPERIMENTS (INTERMOLECULAR RESTRAINTS) |
- Sample preparation
Sample preparation
| Sample conditions | Ionic strength: 0.15 M / pH: 5.5 / Pressure: 1 atm / Temperature: 308.00 K |
|---|---|
| Crystal grow | *PLUS Method: other / Details: NMR |
-NMR measurement
| NMR spectrometer | Type: Bruker AMX 500 / Manufacturer: Bruker / Model: AMX 500 / Field strength: 500 MHz |
|---|
- Processing
Processing
| Software | Name:  AMBER / Classification: refinement AMBER / Classification: refinement | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR software |
| ||||||||||||
| Refinement | Method: RESTRAINED MOLECULAR DYNAMICS / Software ordinal: 1 Details: INITIAL COORDINATES FOR IL-8 WERE TAKEN FROM PDB ENTRY 1IL8; A LINEAR CHAIN FOR THE CXCR-1 FRAGMENT WAS BUILT IN INSIGHT (MSI). THE CXCR-1 FRAGMENT WAS POSITIONED RANDOMLY WITH RESPECT TO ...Details: INITIAL COORDINATES FOR IL-8 WERE TAKEN FROM PDB ENTRY 1IL8; A LINEAR CHAIN FOR THE CXCR-1 FRAGMENT WAS BUILT IN INSIGHT (MSI). THE CXCR-1 FRAGMENT WAS POSITIONED RANDOMLY WITH RESPECT TO IL8 - OBTAIN 40 STARTING CONFORMATIONS. THE INITIAL STRUCTURES WERE THEN REFINED USING RMD WITH THE AMBER ALL ATOM FORCE FIELD AS IMPLIMENTED WITHIN DISCOVER. ALL OF IL8 MONOMER B AND PARTS OF IL8 MONOMER A (2-7, 22-38 AND 51-72) WERE KEPT FIXED DURING THE REFINEMENT SINCE CHEMICAL SHIFT CHANGES INDICATED THAT THESE PORTION OF THE MOLECULE WERE NOT PERTURBED BY PEPTIDE BINDING. THIS MODEL IS THE RESULT OF RESTRAINED ENERGY MINIMIZATION OF THE GEOMETRIC MEAN OF 20 STRUCTURES FROM THE ENSEMBLE (ENTRY 1ILP) SEE JRNL ENTRY FOR MORE DETAILS. | ||||||||||||
| NMR ensemble | Conformer selection criteria: LEAST RESTRAINT VIOLATION ENERGY Conformers calculated total number: 40 / Conformers submitted total number: 1 |
 Movie
Movie Controller
Controller












 PDBj
PDBj













